LRH-1/NR5A2 targets mitochondrial dynamics to reprogram type 1 diabetes macrophages and dendritic cells into an immune tolerance phenotype
- PMID: 39702941
- PMCID: PMC11659195
- DOI: 10.1002/ctm2.70134
LRH-1/NR5A2 targets mitochondrial dynamics to reprogram type 1 diabetes macrophages and dendritic cells into an immune tolerance phenotype
Abstract
Background: The complex aetiology of type 1 diabetes (T1D), characterised by a detrimental cross-talk between the immune system and insulin-producing beta cells, has hindered the development of effective disease-modifying therapies. The discovery that the pharmacological activation of LRH-1/NR5A2 can reverse hyperglycaemia in mouse models of T1D by attenuating the autoimmune attack coupled to beta cell survival/regeneration prompted us to investigate whether immune tolerisation could be translated to individuals with T1D by LRH-1/NR5A2 activation and improve islet survival.
Methods: Peripheral blood mononuclear cells (PBMCs) were isolated from individuals with and without T1D and derived into various immune cells, including macrophages and dendritic cells. Cell subpopulations were then treated or not with BL001, a pharmacological agonist of LRH-1/NR5A2, and processed for: (1) Cell surface marker profiling, (2) cytokine secretome profiling, (3) autologous T-cell proliferation, (4) RNAseq and (5) proteomic analysis. BL001-target gene expression levels were confirmed by quantitative PCR. Mitochondrial function was evaluated through the measurement of oxygen consumption rate using a Seahorse XF analyser. Co-cultures of PBMCs and iPSCs-derived islet organoids were performed to assess the impact of BL001 on beta cell viability.
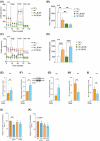
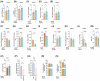
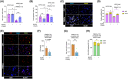
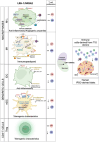
Results: LRH-1/NR5A2 activation induced a genetic and immunometabolic reprogramming of T1D immune cells, marked by reduced pro-inflammatory markers and cytokine secretion, along with enhanced mitohormesis in pro-inflammatory M1 macrophages and mitochondrial turnover in mature dendritic cells. These changes induced a shift from a pro-inflammatory to an anti-inflammatory/tolerogenic state, resulting in the inhibition of CD4+ and CD8+ T-cell proliferation. BL001 treatment also increased CD4+/CD25+/FoxP3+ regulatory T-cells and Th2 cells within PBMCs while decreasing CD8+ T-cell proliferation. Additionally, BL001 alleviated PBMC-induced apoptosis and maintained insulin expression in human iPSC-derived islet organoids.
Conclusion: These findings demonstrate the potential of LRH-1/NR5A2 activation to modulate immune responses and support beta cell viability in T1D, suggesting a new therapeutic approach.
Key points: LRH-1/NR5A2 activation in inflammatory cells of individuals with type 1 diabetes (T1D) reduces pro-inflammatory cell surface markers and cytokine release. LRH-1/NR5A2 promotes a mitohormesis-induced immuno-resistant phenotype to pro-inflammatory macrophages. Mature dendritic cells acquire a tolerogenic phenotype via LRH-1/NR5A2-stimulated mitochondria turnover. LRH-1/NR5A2 agonistic activation expands a CD4+/CD25+/FoxP3+ T-cell subpopulation. Pharmacological activation of LRH-1/NR5A2 improves the survival iPSC-islets-like organoids co-cultured with PBMCs from individuals with T1D.
Keywords: autoimmune diseases; drug development; immune tolerance; pancreatic islets.
© 2024 The Author(s). Clinical and Translational Medicine published by John Wiley & Sons Australia, Ltd on behalf of Shanghai Institute of Clinical Bioinformatics.
Conflict of interest statement
Two patents (WO 2011 144725 A2 and WO 2016 156531 A1) related to BL001 have been published of which B. R. G. and N. C. V. are inventors. M. V.‐P. holds a patent that relates to immunotherapy for T1D and is co‐founder and SEO of Ahead Therapeutics S.L., which aims at the clinical translation of immunotherapies for the treatment of autoimmune diseases. The other authors declare no competing interests related to the current study.
Figures




References
-
- Ward ZJ, Yeh JM, Reddy CL, et al. Estimating the total incidence of type 1 diabetes in children and adolescents aged 0–19 years from 1990 to 2050: a global simulation‐based analysis. Lancet Diabetes Endocrinol. 2022;10(12):848‐858. - PubMed
MeSH terms
Substances
Grants and funding
- MCIN/AEI/10.13039/501100011033/Ministerio de Ciencia E Innovación
- DiabetesCERO Foundation
- 2-SRA-2019-837-S-B/Juvenile Diabetes Research Foundation/Breakthrough TD1
- 17-2013-372/Juvenile Diabetes Research Foundation/Breakthrough TD1
- PI-0727-2010/Consejería de Salud y Consumo, Fundación Pública Andaluza Progreso y Salud, Junta de Andalucía
- PI-0001-2020/Consejería de Salud y Consumo, Fundación Pública Andaluza Progreso y Salud, Junta de Andalucía
- NH/NIH HHS/United States
- DOC_00652/Consejería de Economía, Innovación y Ciencia
- P10.CTS.6359/Consejería de Economía, Innovación y Ciencia
- RO1DK133881-01/DK/NIDDK NIH HHS/United States
- R01 DK133881/DK/NIDDK NIH HHS/United States
- RO1DK126444/DK/NIDDK NIH HHS/United States
- FEDER, UE
- 3-SRA-2022-1201-S-B/Juvenile Diabetes Research Foundation/Breakthrough TD1
- U01 DK127786/DK/NIDDK NIH HHS/United States
- 3-SRA-2023-1307-S-B/Juvenile Diabetes Research Foundation/Breakthrough TD1
- BFU2017-83588-P/Ministerio de Ciencia E Innovación
- PID2021-123083NB-I00/Ministerio de Ciencia E Innovación
- Spanish Diabetes Society
- R01 DK126444/DK/NIDDK NIH HHS/United States
- SE/INV/0008/2022/EU- NextGenerationEU, and the Recovery, Transformation and Resilience Plan
LinkOut - more resources
Full Text Sources
Medical
Research Materials
