Potassium/sodium cation carriers robustly upregulate CD20 antigen by targeting MYC, and synergize with anti- CD20 immunotherapies to eliminate malignant B cells
- PMID: 39704178
- PMCID: PMC12208165
- DOI: 10.3324/haematol.2024.285826
Potassium/sodium cation carriers robustly upregulate CD20 antigen by targeting MYC, and synergize with anti- CD20 immunotherapies to eliminate malignant B cells
Abstract
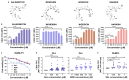
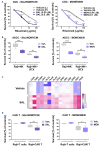
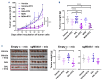
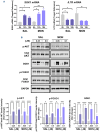
Our investigation uncovers that nanomolar concentrations of salinomycin, monensin, nigericin, and narasin (a group of potassium/ sodium cation carriers) robustly enhance surface expression of CD20 antigen in B-cell-derived tumor cells, including primary malignant cells of chronic lymphocytic leukemia and diffuse large B-cell lymphoma. Experiments in vitro, ex vivo, and animal model reveal a novel approach of combining salinomycin or monensin with therapeutic anti-CD20 monoclonal antibodies or anti-CD20 chimeric antigen receptor T cells, significantly improving non-Hodgkin lymphoma therapy. The results of RNA sequencing, genetic editing, and chemical inhibition delineate the molecular mechanism of CD20 upregulation, at least partially, to the downregulation of MYC, the transcriptional repressor of the MS4A1 gene encoding CD20. Our findings propose the cation carriers as compounds targeting MYC oncogene, which can be combined with anti-CD20 antibodies or adoptive cellular therapies to treat non-Hodgkin lymphoma and mitigate resistance, which frequently depends on the CD20 antigen loss, offering new solutions to improve patient outcomes.
Figures




References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

