scEGOT: single-cell trajectory inference framework based on entropic Gaussian mixture optimal transport
- PMID: 39710672
- PMCID: PMC11665215
- DOI: 10.1186/s12859-024-05988-z
scEGOT: single-cell trajectory inference framework based on entropic Gaussian mixture optimal transport
Abstract
Background: Time-series scRNA-seq data have opened a door to elucidate cell differentiation, and in this context, the optimal transport theory has been attracting much attention. However, there remain critical issues in interpretability and computational cost.
Results: We present scEGOT, a comprehensive framework for single-cell trajectory inference, as a generative model with high interpretability and low computational cost. Applied to the human primordial germ cell-like cell (PGCLC) induction system, scEGOT identified the PGCLC progenitor population and bifurcation time of segregation. Our analysis shows TFAP2A is insufficient for identifying PGCLC progenitors, requiring NKX1-2. Additionally, MESP1 and GATA6 are also crucial for PGCLC/somatic cell segregation.
Conclusions: These findings shed light on the mechanism that segregates PGCLC from somatic lineages. Notably, not limited to scRNA-seq, scEGOT's versatility can extend to general single-cell data like scATAC-seq, and hence has the potential to revolutionize our understanding of such datasets and, thereby also, developmental biology.
Keywords: Epigenetic landscape; Gaussian mixture model; Optimal transport; Single-cell biology; Trajectory inference.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The experiments involving hPGCLCs induced from hiPSCs were approved by the Institutional Review Board of Kyoto University and were also performed in accordance with the guidelines of the Ministry of Education, Culture, Sports, Science, and Technology (MEXT) of Japan. Consent for publication: It does not include individual data. Competing interests: The authors declare no Conflict of interest.
Figures

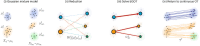




References
-
- Waddington CH. The strategy of the genes: a discussion of some aspects of theoretical biology. Crows Nest: Allen & Unwin; 1957.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

