Targeted dual-receptor phage cocktail against Cronobacter sakazakii: insights into phage-host interactions and resistance mechanisms
- PMID: 39712890
- PMCID: PMC11659082
- DOI: 10.3389/fmicb.2024.1468686
Targeted dual-receptor phage cocktail against Cronobacter sakazakii: insights into phage-host interactions and resistance mechanisms
Abstract
Introduction: Cronobacter sakazakii is a notorious foodborne pathogen, frequently contaminating powdered infant formula and causing life-threatening diseases in infants. The escalating emergence of antibiotics-resistant mutants has led to increased interest in using bacteriophage as an alternative antimicrobial agent.
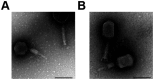
Methods: Two phages, CR8 and S13, were isolated from feces and soil samples and their morphology, physiology, and genomics were characterized. Phage receptor was determined using deletion mutants lacking flgK, rfaC, fhuA, btuB, lamb, or ompC genes, followed by complementation. Phage-resistant mutants were analyzed for phenotypic changes and fitness trade-offs using motility assays and Caco-2 cell invasion models.
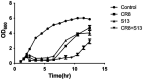
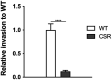
Results: CR8 and S13 were identified as members of Caudoviricetes. Phage CR8 and phage S13 utilize flagella and LPS, respectively, to adhere to host cells. Bacterial challenge assay demonstrated delayed emergence of the resistant mutant as well as stronger lytic activity of a phage cocktail consisting of CR8 and S13 than the single phage treatment. Phenotypic analysis of the phage cocktail resistant strain, designated as CSR strain, revealed that the resistance resulted from the impaired receptor proteins for phage, such as defects in motility and alteration in LPS structure. CSR strain exhibited significant attenuation in invading human intestinal epithelial Caco-2 cells compared to WT cells.
Conclusion: This study demonstrates that the development of the phage cocktail targeting distinct host receptors can serve as a promising antimicrobial strategy to effectively control C. sakazakii.
Keywords: Cronobacter sakazakii; bacteriophage therapy; fitness trade-off; flagella; lipopolysaccharides; phage cocktail; phage resistance.
Copyright © 2024 Kim, Son, Kim, Kim, Nam, Shin and Ryu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures



Similar articles
-
Investigating the biocontrol and anti-biofilm potential of a three phage cocktail against Cronobacter sakazakii in different brands of infant formula.Int J Food Microbiol. 2017 Jul 17;253:1-11. doi: 10.1016/j.ijfoodmicro.2017.04.009. Epub 2017 Apr 21. Int J Food Microbiol. 2017. PMID: 28460269
-
Fitness Trade-Offs in Phage Cocktail-Resistant Salmonella enterica Serovar Enteritidis Results in Increased Antibiotic Susceptibility and Reduced Virulence.Microbiol Spectr. 2022 Oct 26;10(5):e0291422. doi: 10.1128/spectrum.02914-22. Epub 2022 Sep 27. Microbiol Spectr. 2022. PMID: 36165776 Free PMC article.
-
Development of a Bacteriophage Cocktail against Pectobacterium carotovorum Subsp. carotovorum and Its Effects on Pectobacterium Virulence.Appl Environ Microbiol. 2022 Oct 11;88(19):e0076122. doi: 10.1128/aem.00761-22. Epub 2022 Sep 27. Appl Environ Microbiol. 2022. PMID: 36165651 Free PMC article.
-
Trending biocontrol strategies against Cronobacter sakazakii: A recent updated review.Food Res Int. 2020 Nov;137:109385. doi: 10.1016/j.foodres.2020.109385. Epub 2020 Jun 5. Food Res Int. 2020. PMID: 33233087 Review.
-
Fitness Trade-Offs between Phage and Antibiotic Sensitivity in Phage-Resistant Variants: Molecular Action and Insights into Clinical Applications for Phage Therapy.Int J Mol Sci. 2023 Oct 26;24(21):15628. doi: 10.3390/ijms242115628. Int J Mol Sci. 2023. PMID: 37958612 Free PMC article. Review.
References
-
- Calendar R., Abedon S. T. (2005). The bacteriophages. Oxford: Oxford University Press.
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

