This is a preprint.
H3.1K27M-induced misregulation of the TSK/TONSL-H3.1 pathway causes genomic instability
- PMID: 39713323
- PMCID: PMC11661185
- DOI: 10.1101/2024.12.09.627617
H3.1K27M-induced misregulation of the TSK/TONSL-H3.1 pathway causes genomic instability
Update in
-
H3.1K27M-induced misregulation of the TONSOKU-H3.1 pathway causes genomic instability.Nat Commun. 2025 Apr 14;16(1):3547. doi: 10.1038/s41467-025-58892-2. Nat Commun. 2025. PMID: 40229276 Free PMC article.
Abstract
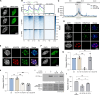
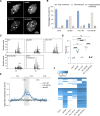
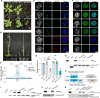
The oncomutation lysine 27-to-methionine in histone H3 (H3K27M) is frequently identified in tumors of patients with diffuse midline glioma-H3K27 altered (DMG-H3K27a). H3K27M inhibits the deposition of the histone mark H3K27me3, which affects the maintenance of transcriptional programs and cell identity. Cells expressing H3K27M are also characterized by defects in genome integrity, but the mechanisms linking expression of the oncohistone to DNA damage remain mostly unknown. In this study, we demonstrate that expression of H3.1K27M in the model plant Arabidopsis thaliana interferes with post-replicative chromatin maturation mediated by the H3.1K27 methyltransferases ATXR5 and ATXR6. As a result, H3.1 variants on nascent chromatin remain unmethylated at K27 (H3.1K27me0), leading to ectopic activity of TONSOKU (TSK), which induces DNA damage and genomic alterations. Elimination of TSK activity suppresses the genome stability defects associated with H3.1K27M expression, while inactivation of specific DNA repair pathways prevents survival of H3.1K27M-expressing plants. Overall, our results suggest that H3.1K27M disrupts the chromatin-based mechanisms regulating TSK/TONSL activity, which causes genomic instability and may contribute to the etiology of DMG-H3K27a.
Conflict of interest statement
Ethics declarations The authors declare no competing interests.
Figures


Similar articles
-
H3.1K27M-induced misregulation of the TONSOKU-H3.1 pathway causes genomic instability.Nat Commun. 2025 Apr 14;16(1):3547. doi: 10.1038/s41467-025-58892-2. Nat Commun. 2025. PMID: 40229276 Free PMC article.
-
Histone H3K27 methylation modulates the dynamics of FANCD2 on chromatin to facilitate NHEJ and genome stability.J Cell Sci. 2018 Jun 21;131(12):jcs215525. doi: 10.1242/jcs.215525. J Cell Sci. 2018. PMID: 29760279
-
The histone H3.1 variant regulates TONSOKU-mediated DNA repair during replication.Science. 2022 Mar 18;375(6586):1281-1286. doi: 10.1126/science.abm5320. Epub 2022 Mar 17. Science. 2022. PMID: 35298257 Free PMC article.
-
Pharmaco-proteogenomic profiling of pediatric diffuse midline glioma to inform future treatment strategies.Oncogene. 2022 Jan;41(4):461-475. doi: 10.1038/s41388-021-02102-y. Epub 2021 Nov 10. Oncogene. 2022. PMID: 34759345 Free PMC article. Review.
-
Histone H3K27M Mutation in Brain Tumors.Adv Exp Med Biol. 2021;1283:43-52. doi: 10.1007/978-981-15-8104-5_3. Adv Exp Med Biol. 2021. PMID: 33155136 Review.
References
-
- Louis D. N. et al., cIMPACT-NOW update 2: diagnostic clarifications for diffuse midline glioma, H3 K27M-mutant and diffuse astrocytoma/anaplastic astrocytoma, IDH-mutant. Acta Neuropathol 135, 639–642 (2018). - PubMed
-
- Schwartzentruber J. et al., Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012). - PubMed
-
- Bender S. et al., Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24, 660–672 (2013). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
