This is a preprint.
Cross-disease modeling of peripheral blood identifies biomarkers of type 2 diabetes predictive of Alzheimer's disease
- PMID: 39713369
- PMCID: PMC11661382
- DOI: 10.1101/2024.12.11.627991
Cross-disease modeling of peripheral blood identifies biomarkers of type 2 diabetes predictive of Alzheimer's disease
Update in
-
Translational disease modeling of peripheral blood identifies type 2 diabetes biomarkers predictive of Alzheimer's disease.NPJ Syst Biol Appl. 2025 May 29;11(1):58. doi: 10.1038/s41540-025-00539-5. NPJ Syst Biol Appl. 2025. PMID: 40442087 Free PMC article.
Abstract
Type 2 diabetes (T2D) is a significant risk factor for Alzheimer's disease (AD). Despite multiple studies reporting this connection, the mechanism by which T2D exacerbates AD is poorly understood. It is challenging to design studies that address co-occurring and comorbid diseases, limiting the number of existing evidence bases. To address this challenge, we expanded the applications of a computational framework called Translatable Components Regression (TransComp-R), initially designed for cross-species translation modeling, to perform cross-disease modeling to identify biological programs of T2D that may exacerbate AD pathology. Using TransComp-R, we combined peripheral blood-derived T2D and AD human transcriptomic data to identify T2D principal components predictive of AD status. Our model revealed genes enriched for biological pathways associated with inflammation, metabolism, and signaling pathways from T2D principal components predictive of AD. The same T2D PC predictive of AD outcomes unveiled sex-based differences across the AD datasets. We performed a gene expression correlational analysis to identify therapeutic hypotheses tailored to the T2D-AD axis. We identified six T2D and two dementia medications that induced gene expression profiles associated with a non-T2D or non-AD state. Finally, we assessed our blood-based T2DxAD biomarker signature in post-mortem human AD and control brain gene expression data from the hippocampus, entorhinal cortex, superior frontal gyrus, and postcentral gyrus. Using partial least squares discriminant analysis, we identified a subset of genes from our cross-disease blood-based biomarker panel that significantly separated AD and control brain samples. Our methodological advance in cross-disease modeling identified biological programs in T2D that may predict the future onset of AD in this population. This, paired with our therapeutic gene expression correlational analysis, also revealed alogliptin, a T2D medication that may help prevent the onset of AD in T2D patients.
Keywords: Alzheimer’s disease; Type 2 diabetes; computational gene correlation analysis; cross-disease modeling; partial least squares discriminant analysis; transcriptomics.
Figures






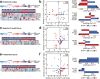
Similar articles
-
Translational disease modeling of peripheral blood identifies type 2 diabetes biomarkers predictive of Alzheimer's disease.NPJ Syst Biol Appl. 2025 May 29;11(1):58. doi: 10.1038/s41540-025-00539-5. NPJ Syst Biol Appl. 2025. PMID: 40442087 Free PMC article.
-
Cross-Species Modeling Identifies Gene Signatures in Type 2 Diabetes Mouse Models Predictive of Inflammatory and Estrogen Signaling Pathways Associated with Alzheimer's Disease Outcomes in Humans.Pac Symp Biocomput. 2025;30:426-440. doi: 10.1142/9789819807024_0031. Pac Symp Biocomput. 2025. PMID: 39670387
-
Mouse-to-human modeling of microglia single-nuclei transcriptomics identifies immune signaling pathways and potential therapeutic candidates associated with Alzheimer's disease.bioRxiv [Preprint]. 2025 Feb 8:2025.02.07.637100. doi: 10.1101/2025.02.07.637100. bioRxiv. 2025. PMID: 39975195 Free PMC article. Preprint.
-
Alzheimer's Disease, Obesity, and Type 2 Diabetes: Focus on Common Neuroglial Dysfunctions (Critical Review and New Data on Human Brain and Models).Brain Sci. 2024 Oct 30;14(11):1101. doi: 10.3390/brainsci14111101. Brain Sci. 2024. PMID: 39595866 Free PMC article. Review.
-
Shared genetic etiology underlying Alzheimer's disease and type 2 diabetes.Mol Aspects Med. 2015 Jun-Oct;43-44:66-76. doi: 10.1016/j.mam.2015.06.006. Epub 2015 Jun 23. Mol Aspects Med. 2015. PMID: 26116273 Free PMC article. Review.
References
-
- Wang K.-C. et al. Risk of Alzheimer’s Disease in Relation to Diabetes: A Population-Based Cohort Study. Neuroepidemiology 38, 237–244 (2012). - PubMed
-
- Janson J. et al. Increased Risk of Type 2 Diabetes in Alzheimer Disease. Diabetes 53, 474–481 (2004). - PubMed
-
- Cheng G., Huang C., Deng H. & Wang H. Diabetes as a risk factor for dementia and mild cognitive impairment: a meta-analysis of longitudinal studies. Internal Medicine Journal 42, 484–491 (2012). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
