Preventing excessive autophagy protects from the pathology of mtDNA mutations in Drosophila melanogaster
- PMID: 39715749
- PMCID: PMC11666730
- DOI: 10.1038/s41467-024-55559-2
Preventing excessive autophagy protects from the pathology of mtDNA mutations in Drosophila melanogaster
Abstract
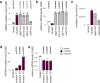
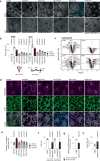
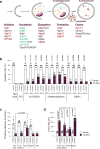
Aberration of mitochondrial function is a shared feature of many human pathologies, characterised by changes in metabolic flux, cellular energetics, morphology, composition, and dynamics of the mitochondrial network. While some of these changes serve as compensatory mechanisms to maintain cellular homeostasis, their chronic activation can permanently affect cellular metabolism and signalling, ultimately impairing cell function. Here, we use a Drosophila melanogaster model expressing a proofreading-deficient mtDNA polymerase (POLγexo-) in a genetic screen to find genes that mitigate the harmful accumulation of mtDNA mutations. We identify critical pathways associated with nutrient sensing, insulin signalling, mitochondrial protein import, and autophagy that can rescue the lethal phenotype of the POLγexo- flies. Rescued flies, hemizygous for dilp1, atg2, tim14 or melted, normalise their autophagic flux and proteasome function and adapt their metabolism. Mutation frequencies remain high with the exception of melted-rescued flies, suggesting that melted may act early in development. Treating POLγexo- larvae with the autophagy activator rapamycin aggravates their lethal phenotype, highlighting that excessive autophagy can significantly contribute to the pathophysiology of mitochondrial diseases. Moreover, we show that the nucleation process of autophagy is a critical target for intervention.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



References
-
- Suomalainen, A. & Battersby, B. J. Mitochondrial diseases: the contribution of organelle stress responses to pathology. Nat. Rev. Mol. Cell Biol.19, 77–92 (2018). - PubMed
-
- Stewart, J. B. & Chinnery, P. F. Extreme heterogeneity of human mitochondrial DNA from organelles to populations. Nat. Rev. Genet22, 106–118 (2020). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

