RcsF-independent mechanisms of signaling within the Rcs phosphorelay
- PMID: 39724052
- PMCID: PMC11709261
- DOI: 10.1371/journal.pgen.1011408
RcsF-independent mechanisms of signaling within the Rcs phosphorelay
Abstract
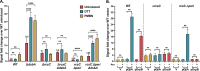
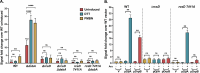
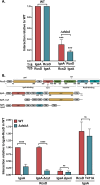
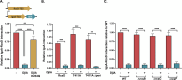
The Rcs (regulator of capsule synthesis) phosphorelay is a conserved cell envelope stress response mechanism in enterobacteria. It responds to perturbations at the cell surface and the peptidoglycan layer from a variety of sources, including antimicrobial peptides, beta-lactams, and changes in osmolarity. RcsF, an outer membrane lipoprotein, is the sensor for this pathway and activates the phosphorelay by interacting with an inner membrane protein IgaA. IgaA is essential; it negatively regulates the signaling by interacting with the phosphotransferase RcsD. We previously showed that RcsF-dependent signaling does not require the periplasmic domain of the histidine kinase RcsC and identified a dominant negative mutant of RcsD that can block signaling via increased interactions with IgaA. However, how the inducing signals are sensed and how signal is transduced to activate the transcription of the Rcs regulon remains unclear. In this study, we investigated how the Rcs cascade functions without its only known sensor, RcsF, and characterized the underlying mechanisms for three distinct RcsF-independent inducers. Previous reports showed that Rcs activity can be induced in the absence of RcsF by a loss of function mutation in the periplasmic oxidoreductase DsbA or by overexpression of the DnaK cochaperone DjlA. We identified an inner membrane protein, DrpB, as a multicopy RcsF-independent Rcs activator in E. coli. The loss of the periplasmic oxidoreductase DsbA and the overexpression of the DnaK cochaperone DjlA each trigger the Rcs cascade in the absence of RcsF by weakening IgaA-RcsD interactions in different ways. In contrast, the cell-division associated protein DrpB uniquely requires the RcsC periplasmic domain for activation; this domain is not needed for RcsF-dependent signaling. This suggests the possibility that the RcsC periplasmic domain acts as a sensor for some Rcs signals. Overall, the results add new understanding to how this complex phosphorelay can be activated by diverse mechanisms.
Copyright: This is an open access article, free of all copyright, and may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose. The work is made available under the Creative Commons CC0 public domain dedication.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Update of
-
RcsF-independent mechanisms of signaling within the Rcs Phosphorelay.bioRxiv [Preprint]. 2024 Sep 3:2024.08.29.610257. doi: 10.1101/2024.08.29.610257. bioRxiv. 2024. Update in: PLoS Genet. 2024 Dec 26;20(12):e1011408. doi: 10.1371/journal.pgen.1011408. PMID: 39372736 Free PMC article. Updated. Preprint.
References
-
- Rogov VV, Rogova NY, Bernhard F, Löhr F, Dötsch V. A disulfide bridge network within the soluble periplasmic domain determines structure and function of the outer membrane protein RCSF. J Biol Chem. 2011;286(21):18775–83. Epub 20110406. doi: 10.1074/jbc.M111.230185 ; PubMed Central PMCID: PMC3099694. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

