Mechanisms of Resistance to KRAS Inhibitors: Cancer Cells' Strategic Use of Normal Cellular Mechanisms to Adapt
- PMID: 39726416
- PMCID: PMC11875783
- DOI: 10.1111/cas.16441
Mechanisms of Resistance to KRAS Inhibitors: Cancer Cells' Strategic Use of Normal Cellular Mechanisms to Adapt
Abstract
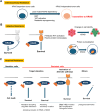
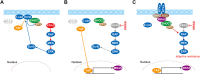
KRAS was long deemed undruggable until the discovery of the switch-II pocket facilitated the development of specific KRAS inhibitors. Despite their introduction into clinical practice, resistance mechanisms can limit their effectiveness. Initially, tumors rely on mutant KRAS, but as they progress, they may shift to alternative pathways, resulting in intrinsic resistance. This resistance can stem from mechanisms like epithelial-to-mesenchymal transition (EMT), YAP activation, or KEAP1 mutations. KRAS inhibition often triggers cellular rewiring to counteract therapeutic pressure. For instance, feedback reactivation of signaling pathways such as MAPK, mediated by receptor tyrosine kinases, supports tumor cell survival. Inhibiting KRAS disrupts protein homeostasis, but reactivation of MAPK or AKT can restore it, aiding tumor cell survival. KRAS inhibition also causes metabolic reprogramming and protein re-localization. The re-localization of E-cadherin and Scribble from the membrane to the cytosol causes YAP to translocate to the nucleus, where it drives MRAS transcription, leading to MAPK reactivation. Emerging evidence indicates that changes in cell identity, such as mucinous differentiation, shifts from alveolar type 2 to type 1 cells, or lineage switching from adenocarcinoma to squamous cell carcinoma, also contribute to resistance. In addition to these nongenetic mechanisms, secondary mutations in KRAS or alterations in upstream/downstream signaling proteins can cause acquired resistance. Secondary mutations in the switch-II pocket disrupt drug binding, and known oncogenic mutations affect drug efficacy. Overcoming these resistance mechanisms involves enhancing the efficacy of drugs targeting mutant KRAS, developing broad-spectrum inhibitors, combining therapies targeting multiple pathways, and integrating immune checkpoint inhibitors.
Keywords: KRAS; lineage plasticity; nongenetic mechanism; resistance; secondary mutation.
© 2024 The Author(s). Cancer Science published by John Wiley & Sons Australia, Ltd on behalf of Japanese Cancer Association.
Conflict of interest statement
H.E. received honoraria from AMGEN and research funding from Lilly‐LOXO and Astellas. N.T. declares no conflicts of interest. H.E. is an editorial board member of Cancer Science.
Figures



References
-
- Simanshu D. K. and Morrison D. K., “A Structure Is Worth a Thousand Words: New Insights for RAS and RAF Regulation,” Cancer Discovery 12 (2022): 899–912, 10.1158/2159-8290.CD-21-1494. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

