Frequent variations and phylogenetic relationships within the genus Secale identified by ND-FISH according to the genome-wide universal oligonucleotides chromosome probes
- PMID: 39726427
- PMCID: PMC11669505
- DOI: 10.3389/fpls.2024.1501642
Frequent variations and phylogenetic relationships within the genus Secale identified by ND-FISH according to the genome-wide universal oligonucleotides chromosome probes
Abstract
Introduction: Rye (Secale cereale L.) played a very important role in wheat genetic improvement and forage production worldwide. However, since rye is a kind of cross-pollinated plant, high levels of genetic heterozygosity and heterogeneity existed in the genome. Genome-wide variation in repeat sequences is one of the most important reasons for chromosome evolution in rye. High-precision cytological identification can effectively identify the heterochromatin or repeat sequence variations in the rye genome, and the relationship between different rye varieties can be identified while obtaining the FISH-karyotype of different rye varieties. The evolution of rye chromosomes can be analyzed by the variation degree of different probes on rye chromosomes.
Methods: All materials were identified by non-denaturing fluorescence in situ hybridization (ND-FISH). Five probes, (AAC)6, Oligo-pSc119.2-1, Oligo-pTa71A-2, Oligo-pSc200, and Oligo-pSc250 were used to identify rye chromosomes.
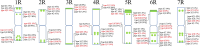
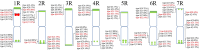
Results: 15 rye varieties including S. cereale (cultivated rye and weedy rye), S. strictum (wild rye), S. sylvestre (wild rye), and S. vavilovii (wild rye) were examined by five oligonucleotides probes. 92 signal sites and 2074 signal patterns were observed, suggesting that high polymorphisms exist in the different rye genomes. The karyotypes of 15 rye varieties were obtained, the frequency of different signal types at each signal site was calculated and the model diagrams of probes (AAC)6, Oligo-pSc119.2-1, Oligo-pTa71A-2, Oligo-pSc200 + Oligo-pSc250 were drawn. The results showed that the rate of variation of different chromosomes of rye was not consistent. 1R, 6R, and 7R have higher variation and genetic diversity, while 2R and 3R have lower variation and are more conserved relative to other chromosomes. The results also indicated that S. sylvestre has a far genetic distance from other rye species, and S. vavilovii might be one of the ancestors of Chinese rye varieties.
Discussion: Results from this study confirmed rapid chromosome change and high levels of chromosome diversity in rye.
Keywords: FISH; evolution; genetic diversity; oligonucleotides; rye.
Copyright © 2024 Li, Sun and Ren.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



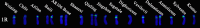

References
-
- An D. G., Ma P. T., Zheng Q., Fu S. L., Li L. H., Han F. P., et al. (2019). Development and molecular cytogenetic identification of a new wheat-rye 4R chromosome disomic addition line with resistances to powdery mildew, stripe rust and sharp eyespot. Theor. Appl. Genet. 132, 257–272. doi: 10.1007/s00122-018-3214-3 - DOI - PubMed
-
- Badaeva E. D., Ruban A. S., Zoshchuk S. A., Surzhikov S. A., Knupffer H., Kilian B. (2016). Molecular cytogenetic characterization of Triticum timopheevii chromosomes provides new insight on genome evolution of T. zhukovskyi . Plant Syst. Evol. 302, 943–956. doi: 10.1007/s00606-016-1309-3 - DOI
LinkOut - more resources
Full Text Sources

