Single-cell analysis identified key macrophage subpopulations associated with atherosclerosis
- PMID: 39726810
- PMCID: PMC11669903
- DOI: 10.1515/med-2024-1088
Single-cell analysis identified key macrophage subpopulations associated with atherosclerosis
Abstract
Background: Atherosclerosis is a lipid-driven inflammatory disease characterized by plaque formation in major arteries. These plaques contain lipid-rich macrophages that accumulate through monocyte recruitment, local macrophage differentiation, and proliferation.
Objective: We identify the macrophage subsets that are closely related to atherosclerosis and reveal the key pathways in the progression of atherosclerotic disease.
Materials and methods: In this study, we characterize the single-cell landscape of atherosclerosis, identifying macrophage subsets closely related to the disease and revealing key pathways in its progression. Using analytical methods like CytoTRACE, Monocle2, Slingshot, and CellChat, we study macrophage differentiation and infer cell trajectory.
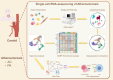
Results: The 8,417 macrophages were divided into six subtypes, macrophages: C0 C1QC+ macrophages, C1 SPP1+ macrophages, C2 FCN1+ macrophages, C3 IGKC+ macrophages, C4 FCER1A+ macrophages, C5CALD1+ macrophages. The results of gene set enrichment analysis, Monocle2, and Slingshot suggest that C2 FCN1+ macrophages may play an important role in the progression of atherosclerosis. C2 FCN1+ macrophages interact with endothelial cells via CCL, CXCL, APP, and other pathways to regulate the progression of atherosclerosis.
Conclusion: We identify a key macrophage subgroup (C2 FCN1+ macrophages) associated with atherosclerosis, which interacts with endothelial cells via CCL, CXCL, APP, and other pathways to regulate disease progression.
Keywords: CCL; FCN1; atherosclerosis; clinical outcome; macrophage.
© 2024 the author(s), published by De Gruyter.
Conflict of interest statement
Conflict of interest: The authors state no conflict of interest.
Figures








Similar articles
-
Unveiling Atherosclerotic Plaque Heterogeneity and SPP1+/VCAN+ Macrophage Subtype Prognostic Significance Through Integrative Single-Cell and Bulk-Seq Analysis.J Inflamm Res. 2024 Apr 23;17:2399-2426. doi: 10.2147/JIR.S454505. eCollection 2024. J Inflamm Res. 2024. PMID: 38681071 Free PMC article.
-
Single-cell and spatial analysis reveals the interaction between ITLN1+ foam cells and SPP1+ macrophages in atherosclerosis.Front Cardiovasc Med. 2025 Feb 13;12:1510082. doi: 10.3389/fcvm.2025.1510082. eCollection 2025. Front Cardiovasc Med. 2025. PMID: 40017519 Free PMC article.
-
Macrophage polarisation associated with atherosclerosis differentially affects their capacity to handle lipids.Atherosclerosis. 2020 Jul;305:10-18. doi: 10.1016/j.atherosclerosis.2020.05.003. Epub 2020 Jun 10. Atherosclerosis. 2020. PMID: 32592946
-
Monocytes, Macrophages, and Metabolic Disease in Atherosclerosis.Front Pharmacol. 2019 Jun 13;10:666. doi: 10.3389/fphar.2019.00666. eCollection 2019. Front Pharmacol. 2019. PMID: 31249530 Free PMC article. Review.
-
Macrophage Plasticity and Atherosclerosis Therapy.Front Mol Biosci. 2021 May 7;8:679797. doi: 10.3389/fmolb.2021.679797. eCollection 2021. Front Mol Biosci. 2021. PMID: 34026849 Free PMC article. Review.
Cited by
-
Single-cell atlas of endothelial cells in atherosclerosis: identifying C1 CXCL12+ ECs as key proliferative drivers for immunological precision therapeutics in atherosclerosis.Front Immunol. 2025 May 12;16:1569988. doi: 10.3389/fimmu.2025.1569988. eCollection 2025. Front Immunol. 2025. PMID: 40421026 Free PMC article.
References
-
- Fan J, Watanabe T. Atherosclerosis: known and unknown. Pathol Int. 2022 Mar;72(3):151–60. 10.1111/pin.13202. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed.... - DOI - PubMed
-
- Herrington W, Lacey B, Sherliker P, Armitage J, Lewington S. Epidemiology of atherosclerosis and the potential to reduce the global burden of atherothrombotic disease. Circ Res. 2016 Feb;118(4):535–46. 10.1161/CIRCRESAHA.115.307611. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed.... - DOI - PubMed
-
- Li Q, Qi L, Zhao K, Ke W, Li T, Xia L. Integrative quantitative and qualitative analysis for the quality evaluation and monitoring of Danshen medicines from different sources using HPLC-DAD and NIR combined with chemometrics. Front Plant Sci. 2022;13:932855. 10.3389/fpls.2022.932855. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed.... - DOI - PMC - PubMed
-
- Williams JW, Winkels H, Durant CP, Zaitsev K, Ghosheh Y, Ley K. Single cell rna sequencing in atherosclerosis research. Circ Res. 2020 Apr;126(9):1112–26. 10.1161/CIRCRESAHA.119.315940. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed.... - DOI - PMC - PubMed
-
- Wang HM, Yang Z, Qi Y, Huang YL, Xiao LX, et al. Addition of risk-enhancing factors improves risk assessment of atherosclerotic cardiovascular disease in middle-aged and older chinese adults: Findings from the chinese multi-provincial cohort study. Cardiovas Innov Appl. 2023;8:27. 10.15212/CVIA.2023.0036. - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
