Insights into polyethylene biodegradative fingerprint of Pseudomonas citronellolis E5 and Rhodococcus erythropolis D4 by phenotypic and genome-based comparative analyses
- PMID: 39726982
- PMCID: PMC11669507
- DOI: 10.3389/fbioe.2024.1472309
Insights into polyethylene biodegradative fingerprint of Pseudomonas citronellolis E5 and Rhodococcus erythropolis D4 by phenotypic and genome-based comparative analyses
Abstract
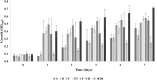
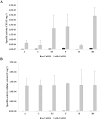
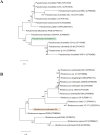
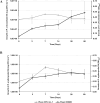
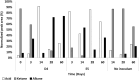
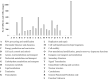
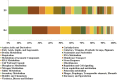
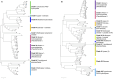
Polyethylene (PE) is the most-produced polyolefin, and consequently, it is the most widely found plastic waste worldwide. PE biodegradation is under study by applying different (micro)organisms in order to understand the biodegradative mechanism in the majority of microbes. This study aims to identify novel bacterial species with compelling metabolic potential and strategic genetic repertoires for PE biodegradation. Pseudomonas citronellolis E5 is newly isolated from solid organic waste contaminated with plastic debris, and Rhodococcus erythropolis D4 was selected for its promising potential in biodegradable plastic determined by its genetic repertoire. P. citronellolis E5 was selected for its ability to grow on PE as the only carbon and energy source. Meaningful extracellular secreted laccase activity was also characterized for D4 during growth on PE (E5 and D4 strains have a laccase activity of (2 ± 1)×10-3 U mg-1 and (3 ± 1)×10-3 U mg-1, respectively). Despite the highest level of cell numbers recorded at 7 days of growth on PE for both strains, the patterns of the metabolic products obtained and degraded during 60 days on PE were dissimilar in the two bacteria at different sampling times. However, they mainly produced metabolites belonging to carboxylic acids and alkanes with varying numbers of carbons in the aliphatic chains. Whole-genome sequence analyses of P. citronellolis E5 compared to R. erythropolis D4 and genetic determinant prediction (by gene annotation and multiple sequence alignment with reference gene products) have been performed, providing a list of 16 and 42 gene products putatively related to different metabolic steps of PE biodegradation. Altogether, these results support insights into PE biodegradation by bacteria of the Pseudomonas and Rhodococcus genera from metabolic and genetic perspectives as a base to build up novel biotechnological platforms.
Keywords: Pseudomonas citronellolis; Rhodococcus erythropolis; gene clusters; genome analysis; laccase activity; polyethylene biodegradation.
Copyright © 2024 Zampolli, Collina, Lasagni and Di Gennaro.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Alvarez H. M. (2019). Biology of Rhodococcus . Cham: Springer Nature. 10.1007/978-3-030-11461-9 - DOI
LinkOut - more resources
Full Text Sources

