Enhanced antibody response to the conformational non-RBD region via DNA prime-protein boost elicits broad cross-neutralization against SARS-CoV-2 variants
- PMID: 39727342
- PMCID: PMC11878195
- DOI: 10.1080/22221751.2024.2447615
Enhanced antibody response to the conformational non-RBD region via DNA prime-protein boost elicits broad cross-neutralization against SARS-CoV-2 variants
Abstract
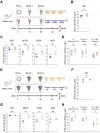
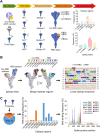
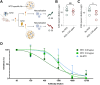
Preventing immune escape of SARS-CoV-2 variants is crucial in vaccine development to ensure broad protection against the virus. Conformational epitopes beyond the RBD region are vital components of the spike protein but have received limited attention in the development of broadly protective SARS-CoV-2 vaccines. In this study, we used a DNA prime-protein boost regimen to evaluate the broad cross-neutralization potential of immune response targeting conformational non-RBD region against SARS-CoV-2 viruses in mice. Mice with enhanced antibody responses targeting conformational non-RBD region show better performance in cross-neutralization against the Wuhan-01, Delta, and Omicron subvariants. Via analyzing the distribution of conformational epitopes, and quantifying epitope-specific binding antibodies, we verified a positive correlation between the proportion of binding antibodies against the N-terminal domain (NTD) supersite (a conformational non-RBD epitope) and SARS-CoV-2 neutralization potency. The current work highlights the importance of high ratio of conformational non-RBD-specific binding antibodies in mediating viral cross-neutralization and provides new insight into overcoming the immune escape of SARS-CoV-2 variants.
Keywords: Conformational non-RBD epitopes; DNA prime-protein boost; NTD supersite-specific binding antibody; SARS-CoV-2; crossneutralization.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures


References
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous
