Identification and Characterization of Two Aryl Sulfotransferases from Deep-Sea Marine Fungi and Their Implications in the Sulfation of Secondary Metabolites
- PMID: 39728146
- PMCID: PMC11677658
- DOI: 10.3390/md22120572
Identification and Characterization of Two Aryl Sulfotransferases from Deep-Sea Marine Fungi and Their Implications in the Sulfation of Secondary Metabolites
Abstract
Sulfation plays a critical role in the biosynthesis of small molecules, regulatory mechanisms such as hormone signaling, and detoxification processes (phase II enzymes). The sulfation reaction is catalyzed by a broad family of enzymes known as sulfotransferases (SULTs), which have been extensively studied in animals due to their medical importance, but also in plant key processes. Despite the identification of some sulfated metabolites in fungi, the mechanisms underlying fungal sulfation remain largely unknown. To address this knowledge gap, we conducted a comprehensive search of available genomes, resulting in the identification of 174 putative SULT genes in the Ascomycota phylum. Phylogenetic analysis and structural modeling revealed that these SULTs belong to the aryl sulfotransferase family, and they are divided into two potential distinct clusters of PAPS-dependent SULTs within the fungal kingdom. SULT genes from two marine fungi isolated from deep-sea hydrothermal vents, Hortaea werneckii UBOCC-A-208029 (HwSULT) and Aspergillus sydowii UBOCC-A-108050 SULT (AsSULT), were selected as representatives of each cluster. Recombinant proteins were expressed in Escherichia coli and biochemically characterized. HwSULT demonstrated high and versatile activity, while AsSULT appeared more substrate-specific. Here, HwSULT was used to sulfate the mycotoxin zearalenone, enhancing its cytotoxicity toward healthy feline intestinal cells.
Keywords: cytotoxic activity; enzymatic sulfation; marine fungi; sulfated metabolites; sulfated mycotoxin; sulfotransferases.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
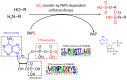







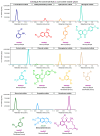
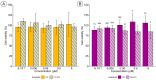
References
-
- Baumann E. Ueber gepaarte Schwefelsäuren im Organismus. Pflüger Arch. 1876;13:285–308. doi: 10.1007/BF01628938. - DOI
-
- Meng L., Shankavaram U., Chen C., Agama K., Fu H., Gonzalez F.J., Weinstein J., Pommier Y. Activation of Aminoflavone (NSC 686288) by a Sulfotransferase Is Required for the Antiproliferative Effect of the Drug and for Induction of Histone Gamma-H2AX. Cancer Res. 2006;66:9656–9664. doi: 10.1158/0008-5472.CAN-06-0796. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

