Interplay of chromatin remodeling BAF complexes in mouse embryonic and epiblast stem cell conversion and maintenance
- PMID: 39730061
- PMCID: PMC11791114
- DOI: 10.1016/j.jbc.2024.108140
Interplay of chromatin remodeling BAF complexes in mouse embryonic and epiblast stem cell conversion and maintenance
Abstract
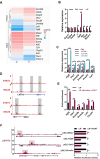
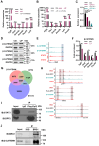
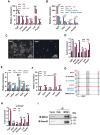
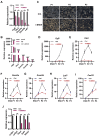
Mouse embryonic stem cells (ESCs) and epiblast stem cells (EpiSCs) are pluripotent stem cells derived from preimplantation and postimplantation embryos, respectively. These cells are capable of interconversion through manipulation of key transcription factors and signaling pathways. While BRG1/BRM-associated factor (BAF) chromatin remodeling complexes are known to play crucial roles in ESC self-renewal and pluripotency, their roles in EpiSCs and their interconversion with ESCs remain unclear. This study demonstrates that the LIF/STAT3 and Wnt signaling pathways, in conjunction with canonical BAF (cBAF) and polycomb repressive complex two complexes, inhibit EpiSC gene expression, thereby preventing ESCs from converting to EpiSCs. Upon removal of LIF, the reduced LIF/STAT3 signaling lifts this inhibition, increasing TGF/nodal pathway activity. Subsequently, the cBAF complex facilitates ESC to EpiSC conversion by promoting EpiSC gene expression. Furthermore, unlike cBAF, inhibition of the ncBAF complex downregulates TGF-β signaling, thereby hindering both ESC to EpiSC conversion and EpiSC maintenance. Moreover, this study revealed the dual mechanisms, methylating histone or non-histone protein STAT3, by which polycomb repressive complex two components participate in the regulation of ESCs to EpiSCs. This research elucidates the interplay between distinct BAF complexes and specific signaling pathways in regulating the conversion and maintenance of ESCs and EpiSCs.
Keywords: BAF complex; ESCs; EpiSCs; LIF/STAT3; TGF-beta.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflict of interest with the contents of the article.
Figures

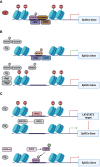
References
-
- Evans M.J., Kaufman M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292:154–156. - PubMed
-
- Brons I.G., Smithers L.E., Trotter M.W., Rugg-Gunn P., Sun B., Chuva de Sousa Lopes S.M., et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature. 2007;448:191–195. - PubMed
-
- Tesar P.J., Chenoweth J.G., Brook F.A., Davies T.J., Evans E.P., Mack D.L., et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature. 2007;448:196–199. - PubMed
-
- Gafni O., Weinberger L., Mansour A.A., Manor Y.S., Chomsky E., Ben-Yosef D., et al. Derivation of novel human ground state naive pluripotent stem cells. Nature. 2013;504:282–286. - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Miscellaneous

