Viral coat proteins decrease the gene silencing activity of cognate and heterologous viral suppressors
- PMID: 39730715
- PMCID: PMC11681101
- DOI: 10.1038/s41598-024-81998-4
Viral coat proteins decrease the gene silencing activity of cognate and heterologous viral suppressors
Abstract
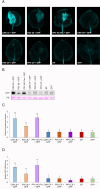
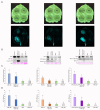
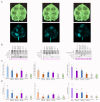
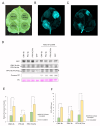
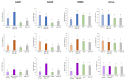
Plant viruses have evolved different viral suppressors of RNA silencing (VSRs) to counteract RNA silencing which is a small RNA-mediated sequence-specific RNA degradation mechanism. Previous studies have already shown that the coat protein (CP) of cucumber mosaic virus (CMV) reduced RNA silencing suppression (RSS) activity of the VSR of CMV, the 2b protein. To demonstrate the universality of this CP-VSR interference, our study included three different viruses: CMV and peanut stunt virus (PSV) from the Bromoviridae, and plum pox virus (PPV) from the Potyviridae family. The RSS activity of the three VSRs (CMV 2b, PSV 2b, and PPV HC-Pro) was compared using Agrobacterium-mediated transient expression in Nicotiana benthamiana and the effect of CMV CP, PSV CP and PPV CP was validated on the RSS activity of their cognate and heterologous VSRs as well. Furthermore, the VSRs were also evaluated in PTGS suppressor-deficient CMV mutant (CMV NVE/10-12/AAA) virus-infected plants. The joint presence of CPs and VSRs resulted in decreased RSS activity in each combination, regardless of the origin of the two proteins, suggesting a universal role of the viral CPs in fine tuning of RSS. Interestingly the PSV CP elicited the strongest negative effect on the RSS activity of all three VSRs.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures
References
-
- Waterhouse, P. M., Wang, M.-B. & Lough, T. Gene silencing as an adaptive defence against viruses. Nature411, 834–842 (2001). - PubMed
-
- Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature431, 343–349 (2004). - PubMed
-
- Vance, V. & Vaucheret, H. RNA silencing in plants-defense and counterdefense. Science292, 2277–2280 (2001). - PubMed
-
- Li, W. X. & Ding, S. W. Viral suppressors of RNA silencing. Curr. Opin. Biotechnol.12, 150–154 (2001). - PubMed
-
- Burgyán, J. & Havelda, Z. Viral suppressors of RNA silencing. Trends Plant Sci.16, 265–272 (2011). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

