Genome-wide analysis of alternative splicing differences in hepatic ischemia reperfusion injury
- PMID: 39732885
- PMCID: PMC11682299
- DOI: 10.1038/s41598-024-82846-1
Genome-wide analysis of alternative splicing differences in hepatic ischemia reperfusion injury
Abstract
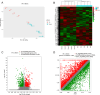
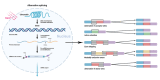
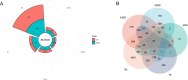
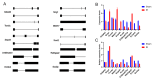
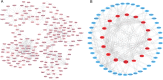
Alternative splicing (AS) contributes to transcript and protein diversity, affecting their structure and function. However, the specific transcriptional regulatory mechanisms underlying AS in the context of hepatic ischemia reperfusion (IR) injury in mice have not been extensively characterized. In this study, we investigated differentially alternatively spliced (DAS) genes and differentially expressed transcripts (DETs) in a mouse model of hepatic IR injury using the high throughput RNA sequencing (RNA-seq) analysis and replicate multivariate analysis of transcript splicing (rMATS) analysis. We further conducted Gene ontology (GO) term enrichment, the Kyoto Encyclopedia of Genes and Genomes (KEGG) database and the protein-protein interaction (PPI) network. A total of 898 DAS genes (p ≤ 0.05) were screened out in the hepatic IR group compared to the sham group, while functional enrichment analysis revealed that DETs and DAS genes were significantly associated with the ATP-dependent chromain, splicesome and metabolic pathways. The expression level of the DAS genes: Gabpb2, Smg1, Tnrc6c, Mettl17, Smpd4, Kcnt2, D16Ertd472e, Rab3gap2, Echdc2 and Ssx2ip were verified by RT-PCR and qRT-PCR. Our findings provide a comprehensive genome-wide view of AS events in hepatic IR injury in mice, enhancing our understanding of AS dynamics and the molecular mechanisms governing alternative pre-mRNA splicing.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethical approval: All procedures in the present study were approved by the Ethics Committee of The First Affiliated Hospital of Harbin Medical University (approval number: No. 2021087), and were also conducted in accordance with the Guide for the Care and Use of Laboratory Animals and with the ARRIVE guidelines. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
Exosome-related gene identification and diagnostic model construction in hepatic ischemia-reperfusion injury.Sci Rep. 2024 Sep 28;14(1):22450. doi: 10.1038/s41598-024-73441-5. Sci Rep. 2024. PMID: 39341981 Free PMC article.
-
Gene Expression Profiling in Ischemic Postconditioning to Alleviate Mouse Liver Ischemia/Reperfusion Injury.Int J Med Sci. 2019 Jan 24;16(2):343-354. doi: 10.7150/ijms.29393. eCollection 2019. Int J Med Sci. 2019. PMID: 30745817 Free PMC article.
-
Identification and functional analysis of differentially expressed genes associated with cerebral ischemia/reperfusion injury through bioinformatics methods.Mol Med Rep. 2018 Aug;18(2):1513-1523. doi: 10.3892/mmr.2018.9135. Epub 2018 Jun 6. Mol Med Rep. 2018. PMID: 29901134 Free PMC article.
-
Differentially expressed genes induced by β-caryophyllene in a rat model of cerebral ischemia-reperfusion injury.Life Sci. 2021 May 15;273:119293. doi: 10.1016/j.lfs.2021.119293. Epub 2021 Mar 8. Life Sci. 2021. PMID: 33705733
-
Deciphering the role of alternative splicing as a potential regulator in fat-tail development of sheep: a comprehensive RNA-seq based study.Sci Rep. 2024 Jan 29;14(1):2361. doi: 10.1038/s41598-024-52855-1. Sci Rep. 2024. PMID: 38287039 Free PMC article. Review.
References
-
- Yang, Y. et al. Hepatocyte-derived Manf alleviates hepatic ischaemia-reperfusion Injury Via regulating endoplasmic reticulum stress-Induced apoptosis in mice. Liver Int.41, 623–639 (2021). - PubMed
Publication types
MeSH terms
Grants and funding
- 81100305, 81470876, 81270527 and 82370643/the National Scientific Foundation of China
- LC2018037/Natural Science Foundation of Heilongjiang Province of China
- LBH-Z11066/Heilongjiang Postdoctoral Foundation
- 2019L01, HYD2020JQ0007/the Scientific Foundation of the First Affiliated Hospital of Harbin Medical University
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

