Biomarker potential of plasma cell-free DNA for cholangiocarcinoma
- PMID: 39735621
- PMCID: PMC11681853
- DOI: 10.1016/j.heliyon.2024.e41008
Biomarker potential of plasma cell-free DNA for cholangiocarcinoma
Abstract
Background: To prevent the development of cholangiocarcinoma, an effective screening opisthorchiasis viverrini and/or differential diagnosis of and the cholangiocarcinoma is crucial needed. The level and quality of cfDNA in plasma are being investigated for their potential role as biomarkers in cholangiocarcinoma.
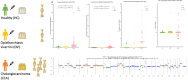
Methods: The study enrolled 43 healthy controls (N), 36 O. viverrini-infected subjects (OV), and 36 cholangiocarcinoma patients (CCA). Plasma cfDNA was quantified by fluorometry (Qubit 4), and qualified analysis, including % tumor fraction, ctDNA, ploidy number, and somatic copy number alteration (SCNA), was conducted using ULP-WGS and analyzed by iChorCNA. The statistical analysis and comparison among the groups were performed.
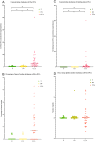
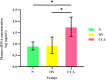
Results: The results showed that cfDNA could effectively differentiate between N and OV from CCA statistically both by DNA amount and quality. Using a cut-off of >20.94 ng/ml, the sensitivity and specificity of the cfDNA concentration were determined to be 86.11% and 98.73% for the differential diagnosis of cholangiocarcinoma, respectively. The ULP-WGS with iChorCNA indicated the % tumor fraction of cfDNA (P < 0.001) and the values of the ploidy number (P < 0.001) of the cholangiocarcinoma group and the other groups were statistically significant. Moreover, the SCNA of the cholangiocarcinoma group was shown to be significantly high in comparison to that of the healthy control group with an odds ratio of 11.688 (P < 0.001).
Conclusion: The use of cfDNA concentration and ULP-WGS for analyzing DNA quality including % tumor fraction, ctDNA concentration, tumor ploidy, and SCNA are useful for the differential diagnosis of cholangiocarcinoma from opisthorchiasis viverrini and healthy individuals.
Keywords: Cell-free DNA; Cholangiocarcinoma; Circulating tumor DNA; Diagnosis; Opisthorchis viverrini; Ultra-low-pass whole-genome sequencing (ULP-WGS); iChorCNA.
© 2024 Chulabhorn International College of Medicine, Thammasat University.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests:Veerachai Thitapakorn reports financial support was provided by 10.13039/501100005790Thammasat University. Veerachai Thitapakorn reports a relationship with Thammasat University that includes: employment. The author declares, there is no conflict of interest for this study. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

Similar articles
-
Investigation of Plasma Cell-Free DNA and MiRNA in Cholangiocarcinoma and Opisthorchiasis Viverrini Patients.Asian Pac J Cancer Prev. 2024 Mar 1;25(3):739-746. doi: 10.31557/APJCP.2024.25.3.739. Asian Pac J Cancer Prev. 2024. PMID: 38546056 Free PMC article.
-
Prognostic value of ultra-low-pass whole-genome sequencing of circulating tumor DNA in hepatocellular carcinoma under systemic treatment.Clin Mol Hepatol. 2024 Apr;30(2):177-190. doi: 10.3350/cmh.2023.0426. Epub 2023 Dec 29. Clin Mol Hepatol. 2024. PMID: 38163441 Free PMC article.
-
Differential plasma proteomes of the patients with Opisthorchiasis viverrini and cholangiocarcinoma identify a polymeric immunoglobulin receptor as a potential biomarker.Heliyon. 2022 Oct 7;8(10):e10965. doi: 10.1016/j.heliyon.2022.e10965. eCollection 2022 Oct. Heliyon. 2022. PMID: 36247154 Free PMC article.
-
Epigenetic Biomarkers and the Wnt/β-Catenin Pathway in Opisthorchis viverrini-associated Cholangiocarcinoma: A Scoping Review on Therapeutic Opportunities.PLoS Negl Trop Dis. 2024 Sep 5;18(9):e0012477. doi: 10.1371/journal.pntd.0012477. eCollection 2024 Sep. PLoS Negl Trop Dis. 2024. PMID: 39236081 Free PMC article.
-
Current State of Knowledge on Blood and Tissue-Based Biomarkers for Opisthorchis viverrini-induced Cholangiocarcinoma: A Review of Prognostic, Predictive, and Diagnostic Markers.Asian Pac J Cancer Prev. 2024 Jan 1;25(1):25-41. doi: 10.31557/APJCP.2024.25.1.25. Asian Pac J Cancer Prev. 2024. PMID: 38285765 Free PMC article. Review.
References
-
- Desai P., Mencia-Trinchant N., Savenkov O., Simon M.S., Cheang G., Lee S., Samuel M., Ritchie E.K., Guzman M.L., Ballman K.V., Roboz G.J., Hassane D.C. Somatic mutations precede acute myeloid leukemia years before diagnosis. Nat. Med. 2018;24:1015–1023. doi: 10.1038/s41591-018-0081-z. - DOI - PMC - PubMed
-
- Panagopoulou M., Karaglani M., Balgkouranidou I., Biziota E., Koukaki T., Karamitrousis E., Nena E., Tsamardinos I., Kolios G., Lianidou E., Kakolyris S., Chatzaki E. Circulating cell-free DNA in breast cancer: size profiling, levels, and methylation patterns lead to prognostic and predictive classifiers. Oncogene. 2019;38:3387–3401. doi: 10.1038/s41388-018-0660-y. - DOI - PubMed
-
- Osumi H., Shinozaki E., Takeda Y., Wakatsuki T., Ichimura T., Saiura A., Yamaguchi K., Takahashi S., Noda T., Zembutsu H. Clinical relevance of circulating tumor DNA assessed through deep sequencing in patients with metastatic colorectal cancer. Cancer Med. 2019;8:408–417. doi: 10.1002/cam4.1913. - DOI - PMC - PubMed
-
- Wang Z., Duan J., Cai S., Han M., Dong H., Zhao J., Zhu B., Wang S., Zhuo M., Sun J., Wang Q., Bai H., Han J., Tian Y., Lu J., Xu T., Zhao X., Wang G., Cao X., Li F., Wang D., Chen Y., Bai Y., Zhao J., Zhao Z., Zhang Y., Xiong L., He J., Gao S., Wang J. Assessment of blood tumor mutational burden as a potential biomarker for immunotherapy in patients with non–small cell lung cancer with use of a next-generation sequencing cancer gene panel. JAMA Oncol. 2019;5:696–702. doi: 10.1001/jamaoncol.2018.7098. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

