Unveiling the molecular landscape of PCOS: identifying hub genes and causal relationships through bioinformatics and Mendelian randomization
- PMID: 39735641
- PMCID: PMC11671271
- DOI: 10.3389/fendo.2024.1431200
Unveiling the molecular landscape of PCOS: identifying hub genes and causal relationships through bioinformatics and Mendelian randomization
Abstract
Background: Polycystic ovary syndrome (PCOS) is a complex endocrine disorder with various contributing factors. Understanding the molecular mechanisms underlying PCOS is essential for developing effective treatments. This study aimed to identify hub genes and investigate potential molecular mechanisms associated with PCOS through a combination of bioinformatics analysis and Mendelian randomization (MR).
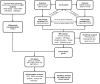
Methods: This study employed bioinformatics analysis in conjunction with MR methods using publicly available databases to identify hub genes. We employed complementary MR methods, including inverse-variance weighted (IVW), to determine the causal relationship between the hub genes and PCOS. Sensitivity analyses were performed to ensure results reliability. Enrichment analysis and immune infiltration analysis were further conducted to assess the role and mechanisms of hub genes in the development of PCOS. Additionally, we validated hub gene expression in both an animal model and serum samples from PCOS patients using qRT-PCR.
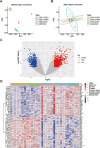
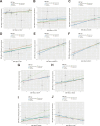
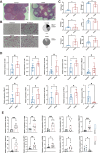
Results: IVW analysis revealed significant associations between 10 hub genes and the risk of PCOS: CD93 [P= 0.004; OR 95%CI= 1.150 (1.046, 1.264)], CYBB [P= 0.013; OR 95%CI= 1.650 (1.113,2.447)], DOCK8 [P= 0.048; OR 95%CI= 1.223 (1.002,1.494)], IRF1 [P= 0.036; OR 95%CI= 1.343 (1.020,1.769)], MBOAT1 [P= 0.033; OR 95%CI= 1.140 (1.011,1.285)], MYO1F [P= 0.012; OR 95%CI= 1.325 (1.065,1.649)], NLRP1 [P= 0.020; OR 95%CI= 1.143 (1.021,1.280)], NOD2 [P= 0.002; OR 95%CI= 1.139 (1.049,1.237)], PIK3R1 [P= 0.040; OR 95%CI= 1.241 (1.010,1.526)], PTER [P= 0.015; OR 95%CI= 0.923 (0.866,0.984)]. No heterogeneity and pleiotropy were observed. Hub genes mainly enriched in positive regulation of cytokine production and TNF signaling pathway, and exhibited positive or negative correlations with different immune cells in individuals with PCOS. qRT-PCR validation in both the rat model and patient serum samples confirmed hub gene expression trends consistent with our combined analysis results.
Conclusions: Our bioinformatics combined with MR analysis revealed that CD93, CYBB, DOCK8, IRF1, MBOAT1, MYO1F, NLRP1, NOD2, PIK3R1 increase the risk of PCOS, while PTER decreases the risk of PCOS. This discovery has implications for clinical decision-making in terms of disease diagnosis, prognosis, treatment strategies, and opens up novel avenues for drug development.
Keywords: Mendelian randomization; SNP; bioinformatic analysis; causal relationship; pcos.
Copyright © 2024 He, Wang, Wang, Deng, Wang, Huang and Lyu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

