Targeted CRISPR/Cas9 Lipid Nanoparticles Elicits Therapeutic Genome Editing in Head and Neck Cancer
- PMID: 39736115
- PMCID: PMC11831472
- DOI: 10.1002/advs.202411032
Targeted CRISPR/Cas9 Lipid Nanoparticles Elicits Therapeutic Genome Editing in Head and Neck Cancer
Abstract
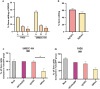
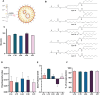
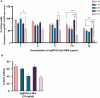
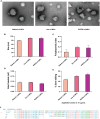
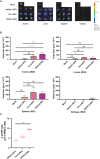
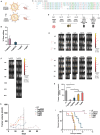
Squamous cell carcinomas of the head and neck (HNSCC) originate in the upper aerodigestive tract, including the oral cavity, pharynx, and larynx. Current treatments of locally advanced HNSCC often lead to high treatment failure, and disease recurrence, resulting in poor survival rates. Advances in mRNA technologies and lipid nanoparticle (LNP) delivery systems led to several clinical trials involving LNP-CRISPR-Cas9 mRNA-based therapeutics. Despite these advances, achieving cell-type-specific extrahepatic mRNA delivery is still challenging. This study introduces a safe and effective intratumoral EGFR-targeted CRISPR-LNP delivery strategy for knocking out SOX2, which is a cancer-specific gene. To assess their therapeutic potential, it is shown that LNPs made from ionizable lipids with helper lipids co-encapsulating Cas9 mRNA and sgRNA targeting SOX2 (sgSOX2), lead to a ≈60% reduction in HNSCC cell viability in vitro. Next, using a xenograft HNSCC mouse model, targeted delivery of 𝜶EGFR- CRISPR-sgSOX2-LNPs to HNSCC cells resulted in a 90% inhibition of tumor growth and a 90% increase in survival for > 84 days, with tumor disappearance observed in 50% of the mice. These findings emphasize the potential of targeted mRNA-Cas9-LNPs in clinically accessible solid tumors, specifically in reaching tumor cells and inducing persistent therapeutic responses in tumors with high-recurrence rates like HNSCC.
Keywords: CRISPR‐Cas9; Genetic Medicines; Head and neck cancer; Lipid nanoparticles; SOX2; mRNA.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
D.P. receives licensing fees (to patents on which he was an inventor) from, invested in, consults (or on scientific advisory boards or boards of directors) for, lectured (and received a fee), or conducts sponsored research at TAU for the following entities: ART Biosciences, BioNtech SE, Earli Inc., Geneditor Biologics Inc.; Kernal Biologics, Newphase Ltd., NeoVac Ltd., RiboX Therapeutics, Roche, Teva Pharmaceuticals Inc. All other authors declare no competing financial interests.
Figures
References
-
- ChowLaura Q., N. Engl. J. Med. 2020, 382, 60.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous
