The evolutionary history of Plasmodium falciparum from mitochondrial and apicoplast genomes of China-Myanmar border isolates
- PMID: 39736695
- PMCID: PMC11686842
- DOI: 10.1186/s13071-024-06629-3
The evolutionary history of Plasmodium falciparum from mitochondrial and apicoplast genomes of China-Myanmar border isolates
Abstract
Background: The frequent communication between African and Southeast Asian (SEA) countries has led to the risk of imported malaria cases in the China-Myanmar border (CMB) region. Therefore, tracing the origins of new malaria infections is important in the maintenance of malaria-free zones in this border region. A new genotyping tool based on a robust mitochondrial (mt) /apicoplast (apico) barcode was developed to estimate genetic diversity and infer the evolutionary history of Plasmodium falciparum across the major distribution ranges. However, the mt/apico genomes of P. falciparum isolates from the CMB region to date are poorly characterized, even though this region is highly endemic to P. falciparum malaria.
Methods: We have sequenced the whole mt/apico genome of 34 CMB field isolates and utilized a published data set of 147 mt/apico genome sequences to present global genetic diversity and to revisit the evolutionary history of the CMB P. falciparum.
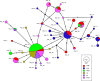
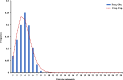
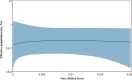
Results: Genetic differentiation based on mt/apico genome of P. falciparum revealed that the CMB (Lazan, Myanmar) isolates presented high genetic diversity with several characteristics of ancestral populations and shared many of the genetic features with West Thailand (Mae Sot; WTH) and to some extent West African (Banjul, Gambia; Navrongo, Ghana; WAF) isolates. The reconstructed haplotype network displayed that the CMB and WTH P. falciparum isolates have the highest representation (five) in the five ancestral (central) haplotypes (H1, H2, H4, H7, and H8), which are comparatively older than isolates from other SEA populations as well as the WAF populations. In addition, the highest estimate of the time to the Most Recent Common Ancestor (TMRCA) of 42,400 (95% CI 18,300-82100) years ago was presented by the CMB P. falciparum compared to the other regional populations. The statistically significant negative values of Fu's Fs with unimodal distribution in pairwise mismatch distribution curves indicate past demographic expansions in CMB P. falciparum with slow population expansion between approximately 12,500-20,000 ybp.
Conclusions: The results on the complete mt/apico genome sequence analysis of the CMB P. falciparum indicated high genetic diversity with ancient population expansion and TMRCA, and it seems probable that P. falciparum might have existed in CMB, WTH, and WAF for a long time before being introduced into other Southeast Asian countries or regions. To reduce the impact of sample size or geographic bias on the estimate of the evolutionary timeline, future studies need to expand the range of sample collection and ensure the representativeness of samples across geographic distributions. Additionally, by mapping global patterns of mt/apico genome polymorphism, we will gain valuable insights into the evolutionary history of P. falciparum and optimised strategies for controlling P. falciparum malaria at international borders.
Keywords: Plasmodium falciparum; Apicoplast; China-Myanmar border; Evolutionary history; Mitochondrial.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The authors assert that all procedures contributing to this work complied with the principles of the Declaration of Helsinki and were approved by the Internal Review Board of Naval Medical University. The patients/participants provided their written informed consent to participate in this study. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures
References
-
- WHO. World Malaria Report 2023. Geneva: World Health Organization; 2023. https://iris.who.int/handle/10665/374472. Accessed 03 May 2024.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

