GATD3A-deficiency-induced mitochondrial dysfunction facilitates senescence of fibroblast-like synoviocytes and osteoarthritis progression
- PMID: 39738099
- PMCID: PMC11685659
- DOI: 10.1038/s41467-024-55335-2
GATD3A-deficiency-induced mitochondrial dysfunction facilitates senescence of fibroblast-like synoviocytes and osteoarthritis progression
Abstract
Accumulating evidence indicates that cellular senescence is closely associated with osteoarthritis. However, there is limited research on the mechanisms underlying fibroblast-like synoviocyte senescence and its impact on osteoarthritis progression. Here, we elucidate a positive correlation between fibroblast-like synoviocyte senescence and osteoarthritis progression and reveal that GATD3A deficiency induces fibroblast-like synoviocyte senescence. Mechanistically, GATD3A deficiency enhances the binding of Sirt3 to MDH2, leading to deacetylation and decreased activity of MDH2. Reduced MDH2 activity impairs tricarboxylic acid cycle flux, resulting in mitochondrial dysfunction and fibroblast-like synoviocyte senescence. Intra-articular injection of recombinant adeno-associated virus carrying GATD3A significantly alleviates the osteoarthritis phenotype in male mice. This study increases our current understanding of GATD3A function. In particular, we reveal a novel mechanism of fibroblast-like synoviocyte senescence, suggesting that targeting GATD3A is a potential therapeutic approach for osteoarthritis.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures
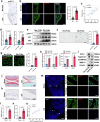







References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

