piRNA processing within non-membrane structures is governed by constituent proteins and their functional motifs
- PMID: 39739617
- PMCID: PMC12138184
- DOI: 10.1111/febs.17360
piRNA processing within non-membrane structures is governed by constituent proteins and their functional motifs
Abstract
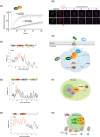
Discovered two decades ago, PIWI-interacting RNAs (piRNAs) are crucial for silencing transposable elements (TEs) in animal gonads, thereby protecting the germline genome from harmful transposition, and ensuring species continuity. Silencing of TEs is achieved through transcriptional and post-transcriptional suppression by piRNAs and the PIWI clade of Argonaute proteins within non-membrane structured organelle. These structures are composed of proteins involved in piRNA processing, including PIWIs and other proteins by distinct functional motifs such as the Tudor domain, LOTUS, and intrinsic disordered regions (IDRs). This review highlights recent advances in understanding the roles of these conserved proteins and structural motifs in piRNA biogenesis. We explore the molecular mechanisms of piRNA biogenesis, with a primary focus on Drosophila as a model organism, identifying common themes and species-specific variations. Additionally, we extend the discussion to the roles of these components in nongonadal tissues.
Keywords: Drosophila germline; Tudor domain‐containing proteins; liquid–liquid phase separation; non‐membrane nuage; piRNAs.
© 2024 The Author(s). The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
The absence of core piRNA biogenesis factors does not impact efficient transposon silencing in Drosophila.PLoS Biol. 2023 Jun 6;21(6):e3002099. doi: 10.1371/journal.pbio.3002099. eCollection 2023 Jun. PLoS Biol. 2023. PMID: 37279192 Free PMC article.
-
The Tudor domain protein Tapas, a homolog of the vertebrate Tdrd7, functions in the piRNA pathway to regulate retrotransposons in germline of Drosophila melanogaster.BMC Biol. 2014 Oct 6;12:61. doi: 10.1186/s12915-014-0061-9. BMC Biol. 2014. PMID: 25287931 Free PMC article.
-
Drosophila Piwi distinguishes transposons from mRNAs by piRNA complementarity and abundance.Cell Rep. 2024 Dec 24;43(12):115020. doi: 10.1016/j.celrep.2024.115020. Epub 2024 Dec 4. Cell Rep. 2024. PMID: 39636727
-
piRNA-Guided Genome Defense: From Biogenesis to Silencing.Annu Rev Genet. 2018 Nov 23;52:131-157. doi: 10.1146/annurev-genet-120417-031441. Annu Rev Genet. 2018. PMID: 30476449 Free PMC article. Review.
-
One Loop to Rule Them All: The Ping-Pong Cycle and piRNA-Guided Silencing.Trends Biochem Sci. 2016 Apr;41(4):324-337. doi: 10.1016/j.tibs.2015.12.008. Epub 2016 Jan 19. Trends Biochem Sci. 2016. PMID: 26810602 Free PMC article. Review.
References
-
- Vagin V, Sigova A, Li C, Seitz H, Gvozdev V & Zamore PD (2006) A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324. - PubMed
-
- Wang X, Ramat A, Simonelig M & Liu MF (2023) Emerging roles and functional mechanisms of PIWI‐interacting RNAs. Nat Rev Mol Cell Biol 24, 123–141. - PubMed
-
- Aravin AA, Naumova NM, Tulin AV, Vagin VV, Rozovsky YM & Gvozdev VA (2001) Double‐stranded RNA‐mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr Biol 11, 1017–1027. - PubMed
-
- Aravin A, Gaidatzis D, Pfeffer S, Lagos‐Quintana M, Landgraf P, Iovino N, Morris P, Brownstein MJ, Kuramochi‐Miyagawa S, Nakano T et al. (2006) A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

