Identification of O-glycosylation related genes and subtypes in ulcerative colitis based on machine learning
- PMID: 39739658
- PMCID: PMC11687659
- DOI: 10.1371/journal.pone.0311495
Identification of O-glycosylation related genes and subtypes in ulcerative colitis based on machine learning
Abstract
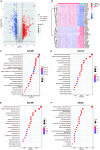
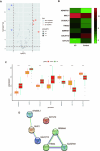
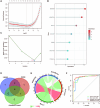
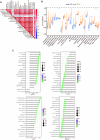
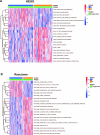
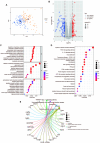
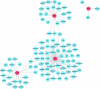
Ulcerative colitis (UC) is an immune-related inflammatory bowel disease, with its underlying mechanisms being a central area of clinical research. O-GlcNAcylation plays a critical role in regulating immunity progression and the occurrence of inflammatory diseases and tumors. Yet, the mechanism of O-GlcNAc-associated colitis remains to be elucidated. To this end, the transcriptional and clinical data of GSE75214 and GSE92415 from the GEO database was hereby examined, and genes MUC1, ADAMTS1, GXYLT2, and SEMA5A were found to be significantly related to O-GlcNAcylation using machine learning methods. Based on the four hub genes, two UC subtypes were built. Notably, subtype B might be prone to developing colitis-associated colorectal cancer (CAC). This study delved into the role of intestinal glycosylation changes, especially the O-GlcNAcylation, and forged a foundation for further research on the occurrence and development of UC. Overall, understanding the role of O-GlcNAcylation in UC could have significant implications for diagnosis and treatment, offering valuable insights into the disease's progression.
Copyright: © 2024 Lu et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Zhang S. Z., Zhao X. H. & Zhang D. C. Cellular and molecular immunopathogenesis of ulcerative colitis. Cell Mol Immunol 3, 35–40 (2006). - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

