Higher-order transient structures and the principle of dynamic connectivity in membrane signaling
- PMID: 39739805
- PMCID: PMC11725812
- DOI: 10.1073/pnas.2421280121
Higher-order transient structures and the principle of dynamic connectivity in membrane signaling
Abstract
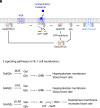
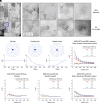
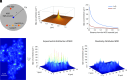
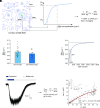
We examine the role of higher-order transient structures (HOTS) in M2R regulation of GIRK channels. Electron microscopic membrane protein location maps show that both proteins form HOTS that exhibit a statistical bias to be near each other. Theoretical calculations and electrophysiological measurements suggest that channel activity is isolated near larger M2R HOTS. By invoking weak interactions that permit transient binding of M2R to M2R and GIRK to GIRK (i-i interactions) and M2R to GIRK (i-j interactions), the distribution patterns and electrophysiological properties of HL-1 cells are replicated in a reaction-diffusion simulation. We propose the principle of dynamic connectivity to explain communication between protein components of a membrane signaling pathway. Dynamic connectivity is mediated by weak, transient interactions between proteins. HOTS created by weak i-i interactions, and statistical biases created by weak i-j interactions promoted by the multivalence of HOTS, are the key elements of dynamic connectivity.
Keywords: GPCR; HOTS; higher-order transient structure; membrane signaling; self-assembly.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures


Similar articles
-
GPCR-dependent biasing of GIRK channel signaling dynamics by RGS6 in mouse sinoatrial nodal cells.Proc Natl Acad Sci U S A. 2020 Jun 23;117(25):14522-14531. doi: 10.1073/pnas.2001270117. Epub 2020 Jun 8. Proc Natl Acad Sci U S A. 2020. PMID: 32513692 Free PMC article.
-
Receptor-mediated inhibition of G protein-coupled inwardly rectifying potassium channels involves G(alpha)q family subunits, phospholipase C, and a readily diffusible messenger.J Biol Chem. 2001 May 18;276(20):16720-30. doi: 10.1074/jbc.M100207200. Epub 2001 Mar 8. J Biol Chem. 2001. PMID: 11279027
-
Agonist-induced localization of Gq-coupled receptors and G protein-gated inwardly rectifying K+ (GIRK) channels to caveolae determines receptor specificity of phosphatidylinositol 4,5-bisphosphate signaling.J Biol Chem. 2010 Dec 31;285(53):41732-9. doi: 10.1074/jbc.M110.153312. Epub 2010 Nov 1. J Biol Chem. 2010. PMID: 21041306 Free PMC article.
-
Structural Insights into GIRK Channel Function.Int Rev Neurobiol. 2015;123:117-60. doi: 10.1016/bs.irn.2015.05.014. Epub 2015 Jun 22. Int Rev Neurobiol. 2015. PMID: 26422984 Review.
-
RGS Redundancy and Implications in GPCR-GIRK Signaling.Int Rev Neurobiol. 2015;123:87-116. doi: 10.1016/bs.irn.2015.05.010. Epub 2015 Jun 22. Int Rev Neurobiol. 2015. PMID: 26422983 Review.
Cited by
-
Higher-order transient membrane protein structures.Proc Natl Acad Sci U S A. 2025 Jan 7;122(1):e2421275121. doi: 10.1073/pnas.2421275121. Epub 2024 Dec 31. Proc Natl Acad Sci U S A. 2025. PMID: 39739811 Free PMC article.
-
Intersection of GPCR trafficking and cAMP signaling at endomembranes.J Cell Biol. 2025 Apr 7;224(4):e202409027. doi: 10.1083/jcb.202409027. Epub 2025 Mar 25. J Cell Biol. 2025. PMID: 40131202 Review.
-
Detergent-free isolation and characterization of amyloid precursor protein C99 in E. coli native lipid-nanodiscs using non-ionic polymer.Protein Sci. 2025 Sep;34(9):e70276. doi: 10.1002/pro.70276. Protein Sci. 2025. PMID: 40828514 Free PMC article.
-
Molecular contacts in self-assembling clusters of membrane proteins.Proc Natl Acad Sci U S A. 2025 Jul;122(26):e2507112122. doi: 10.1073/pnas.2507112122. Epub 2025 Jun 23. Proc Natl Acad Sci U S A. 2025. PMID: 40549920 Free PMC article.
References
-
- Logothetis D. E., Kurachi Y., Galper J., Neer E. J., Clapham D. E., The beta gamma subunits of GTP-binding proteins activate the muscarinic K+ channel in heart. Nature 325, 321–326 (1987). - PubMed
-
- Sakmann B., Noma A., Trautwein W., Acetylcholine activation of single muscarinic K+ channels in isolated pacemaker cells of the mammalian heart. Nature 303, 250–253 (1983). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

