The CYLD-PARP1 feedback loop regulates DNA damage repair and chemosensitivity in breast cancer cells
- PMID: 39739815
- PMCID: PMC11725943
- DOI: 10.1073/pnas.2413890121
The CYLD-PARP1 feedback loop regulates DNA damage repair and chemosensitivity in breast cancer cells
Abstract
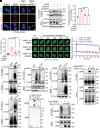
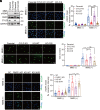
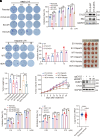
Poly(ADP-ribose) polymerase 1 (PARP1) plays a crucial role in DNA repair and genomic stability maintenance. However, the regulatory mechanisms governing PARP1 activity, particularly through deubiquitination, remain poorly elucidated. Using a deubiquitinase (DUB) library binding screen, we identified cylindromatosis (CYLD) as a bona fide DUB for PARP1 in breast cancer cells. Mechanistically, CYLD is recruited by PARP1 to DNA lesions upon genotoxic stress, where it cleaves K63-linked polyubiquitin chains on PARP1 at residues K748, K940, and K949, resulting in compromised PARP1 activation. In a reciprocal manner, PARP1 PARylates CYLD at sites E191, E231, E259, and E509, thereby enhancing its DUB activity. Consequently, depletion of CYLD leads to increased efficiency in base excision repair and confers breast cancer cells with resistance to alkylating agents. Conversely, overexpression of CYLD enhances sensitivity to PARP inhibitors (PARPi) even in homologous recombination-proficient breast cancer cells. These findings offer unique insights into the intricate interplay between CYLD and PARP1 in DNA repair, underscoring the pivotal role of targeting this regulatory axis for breast cancer chemotherapy.
Keywords: CYLD; PARP1; PARylation; deubiquitination; drug sensitivity.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures



Similar articles
-
The deubiquitination-PARylation positive feedback loop of the USP10-PARP1 axis promotes DNA damage repair and affects therapeutic efficacy of PARP1 inhibitor.Oncogene. 2025 Aug;44(29):2515-2529. doi: 10.1038/s41388-025-03428-7. Epub 2025 May 2. Oncogene. 2025. PMID: 40316740 Free PMC article.
-
DNMT1 is required for efficient DSB repair and maintenance of replication fork stability, and its loss reverses resistance to PARP inhibitors in cancer cells.Oncogene. 2025 Jul;44(27):2283-2302. doi: 10.1038/s41388-025-03409-w. Epub 2025 Apr 15. Oncogene. 2025. PMID: 40234721
-
FGFR3-induced Y158 PARP1 phosphorylation promotes PARP inhibitor resistance via BRG1/MRE11-mediated DNA repair in breast cancer models.J Clin Invest. 2025 May 29;135(14):e173757. doi: 10.1172/JCI173757. eCollection 2025 Jul 15. J Clin Invest. 2025. PMID: 40460005 Free PMC article.
-
"Yin-Yang" of PARP1 in genotoxic and inflammatory response.DNA Repair (Amst). 2025 Aug;152:103858. doi: 10.1016/j.dnarep.2025.103858. Epub 2025 Jun 11. DNA Repair (Amst). 2025. PMID: 40527058 Review.
-
Poly(ADP-ribose) polymerase (PARP) inhibitors for the treatment of ovarian cancer.Cochrane Database Syst Rev. 2022 Feb 16;2(2):CD007929. doi: 10.1002/14651858.CD007929.pub4. Cochrane Database Syst Rev. 2022. PMID: 35170751 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

