Structural and functional dynamics of human cone cGMP-phosphodiesterase important for photopic vision
- PMID: 39739818
- PMCID: PMC11725853
- DOI: 10.1073/pnas.2419732121
Structural and functional dynamics of human cone cGMP-phosphodiesterase important for photopic vision
Erratum in
-
Correction for Singh et al., Structural and functional dynamics of human cone cGMP-phosphodiesterase important for photopic vision.Proc Natl Acad Sci U S A. 2025 Apr 8;122(14):e2505043122. doi: 10.1073/pnas.2505043122. Epub 2025 Mar 28. Proc Natl Acad Sci U S A. 2025. PMID: 40153471 Free PMC article. No abstract available.
Abstract
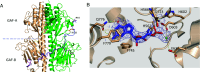
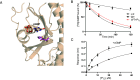
Cone cGMP-phosphodiesterase (PDE6) is the key effector enzyme for daylight vision, and its properties are critical for shaping distinct physiology of cone photoreceptors. We determined the structures of human cone PDE6C in various liganded states by single-particle cryo-EM that reveal essential functional dynamics and adaptations of the enzyme. Our analysis exposed the dynamic nature of PDE6C association with its regulatory γ-subunit (Pγ) which allows openings of the catalytic pocket in the absence of phototransduction signaling, thereby controlling photoreceptor noise and sensitivity. We demonstrate evolutionarily recent adaptations of PDE6C stemming from residue substitutions in the Pγ subunit and the noncatalytic cGMP binding site and influencing the Pγ dynamics in holoPDE6C. Thus, our structural analysis sheds light on the previously unrecognized molecular evolution of the effector enzyme in cones that advances adaptation for photopic vision.
Keywords: cryo-EM; phosphodiesterase 6; photoreceptor; phototransduction.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures



Similar articles
-
cGMP Signaling in Photoreceptor Degeneration.Int J Mol Sci. 2023 Jul 7;24(13):11200. doi: 10.3390/ijms241311200. Int J Mol Sci. 2023. PMID: 37446378 Free PMC article. Review.
-
Probing the mechanism by which the retinal G protein transducin activates its biological effector PDE6.J Biol Chem. 2024 Feb;300(2):105608. doi: 10.1016/j.jbc.2023.105608. Epub 2023 Dec 28. J Biol Chem. 2024. PMID: 38159849 Free PMC article.
-
Exchange of Cone for Rod Phosphodiesterase 6 Catalytic Subunits in Rod Photoreceptors Mimics in Part Features of Light Adaptation.J Neurosci. 2015 Jun 17;35(24):9225-35. doi: 10.1523/JNEUROSCI.3563-14.2015. J Neurosci. 2015. PMID: 26085644 Free PMC article.
-
Reconstitution of the phosphodiesterase 6 maturation process important for photoreceptor cell function.J Biol Chem. 2024 Jan;300(1):105576. doi: 10.1016/j.jbc.2023.105576. Epub 2023 Dec 16. J Biol Chem. 2024. PMID: 38110033 Free PMC article.
-
Photoreceptor phosphodiesterase (PDE6): activation and inactivation mechanisms during visual transduction in rods and cones.Pflugers Arch. 2021 Sep;473(9):1377-1391. doi: 10.1007/s00424-021-02562-x. Epub 2021 Apr 15. Pflugers Arch. 2021. PMID: 33860373 Free PMC article. Review.
Cited by
-
Roles of Cyclic Nucleotide Phosphodiesterases in Signal Transduction Pathways in the Nematode Caenorhabditis elegans.Cells. 2025 Jul 30;14(15):1174. doi: 10.3390/cells14151174. Cells. 2025. PMID: 40801606 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

