Identifying the NEAT1/miR-26b-5p/S100A2 axis as a regulator in Parkinson's disease based on the ferroptosis-related genes
- PMID: 39739972
- PMCID: PMC11687868
- DOI: 10.1371/journal.pone.0316179
Identifying the NEAT1/miR-26b-5p/S100A2 axis as a regulator in Parkinson's disease based on the ferroptosis-related genes
Abstract
Objectives: Parkinson's disease (PD) is a complex neurodegenerative disease with unclear pathogenesis. Some recent studies have shown that there is a close relationship between PD and ferroptosis. We aimed to identify the ferroptosis-related genes (FRGs) and construct competing endogenous RNA (ceRNA) networks to further assess the pathogenesis of PD.
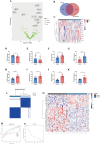
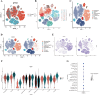
Methods: Expression of 97 substantia nigra (SN) samples were obtained and intersected with FRGs. Bioinformatics analysis, including the gene set enrichment analysis (GSEA), consensus cluster analysis, weight gene co-expression network analysis (WGCNA), and machine learning algorithms, were employed to assess the feasible differentially expressed genes (DEGs). Characteristic signature genes were used to create novel diagnostic models and construct competing endogenous RNA (ceRNA) regulatory network for PD, which were further verified by in vitro experiments and single-cell RNA sequencing (scRNA-seq).
Results: A total of 453 DEGs were identified and 11 FRGs were selected. We sorted the entire PD cohort into two subtypes based on the FRGs and obtained 67 hub genes. According to the five machine algorithms, 4 features (S100A2, GNGT1, NEUROD4, FCN2) were screened and used to create a PD diagnostic model. Corresponding miRNAs and lncRNAs were predicted to construct a ceRNA network. The scRNA-seq and experimental results showed that the signature model had a certain diagnostic effect and lncRNA NEAT1 might regulate the progression of ferroptosis in PD via the NEAT1/miR-26b-5p/S100A2 axis.
Conclusion: The diagnostic signatures based on the four FRGs had certain diagnostic and individual effects. NEAT1/miR-26b-5p/S100A2 axis is associated with ferroptosis in the pathogenesis of PD. Our findings provide new solutions for treating PD.
Copyright: © 2024 Li, Guo. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures






References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

