Efficient genome editing in medaka (Oryzias latipes) using a codon-optimized SaCas9 system
- PMID: 39743295
- PMCID: PMC11693391
- DOI: 10.1631/jzus.B2300899
Efficient genome editing in medaka (Oryzias latipes) using a codon-optimized SaCas9 system
Abstract
The clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) system, belonging to the type II CRISPR/Cas system, is an effective gene-editing tool widely used in different organisms, but the size of Streptococcus pyogenes Cas9 (SpCas9) is quite large (4.3 kb), which is not convenient for vector delivery. In this study, we used a codon-optimized Staphylococcus aureus Cas9 (SaCas9) system to edit the tyrosinase (tyr), oculocutaneous albinism II (oca2), and paired box 6.1 (pax6.1) genes in the fish model medaka(Oryzias latipes), in which the size of SaCas9 (3.3 kb) is much smaller and the necessary protospacer-adjacent motif (PAM) sequence is 5'-NNGRRT-3'. We also used a transfer RNA (tRNA)-single-guide RNA (sgRNA) system to express the functional sgRNA by transcription eitherin vivo or in vitro, and the combination of SaCas9 and tRNA-sgRNA was used to edit the tyr gene in the medaka genome. The SaCas9/sgRNA and SaCas9/tRNA-sgRNA systems were shown to edit the medaka genome effectively, while the PAM sequence is an essential part for the efficiency of editing. Besides, tRNA can improve the flexibility of the system by enabling the sgRNA to be controlled by a common promoter such as cytomegalovirus. Moreover, the all-in-one cassette cytomegalovirus (CMV)-SaCas9-tRNA-sgRNA-tRNA is functional in medaka gene editing. Taken together, the codon-optimized SaCas9 system provides an alternative and smaller tool to edit the medaka genome and potentially other fish genomes.
CRISPR-Cas9系统属于II型CRISPR/Cas系统,作为一种有效的基因编辑工具被广泛应用于不同生物体中,但化脓性链球菌(Streptococcus pyogenes)Cas9(SpCas9)体积较大(4.3 kb),因此使用载体传递较为不便。本研究利用密码子优化后的金黄色葡萄球菌(Staphylococcus aureus)Cas9(SaCas9)系统进行了青鳉(Oryzias latipes)的tyr、oca2和pax6.1基因编辑。SaCas9片段(3.3 kb)要小,所需的原型间隔相邻基序(PAM)序列为5'-NNGRRT-3'。此外,本研究还利用tRNA-sgRNA系统在体内和体外转录表达功能性sgRNA,经SaCas9和tRNA-sgRNA的组合编辑青鳉基因组中的tyr基因。实验结果表明,SaCas9/sgRNA和SaCas9/tRNA-sgRNA系统均能有效编辑青鳉基因组,而PAM序列是该系统进行有效编辑的关键部分。此外,tRNA还能使sgRNA受巨细胞病毒等常见启动子的控制,从而提高系统的适应性。本研究还发现,CMV-SaCas9-tRNA-sgRNA-tRNA一体化结构在青鳉基因编辑中同样能发挥作用。综上,经密码子优化的SaCas9系统为编辑青鳉及潜在的其他鱼类基因组提供了一种更便捷的工具。.
CRISPR-Cas9系统属于II型CRISPR/Cas系统,作为一种有效的基因编辑工具被广泛应用于不同生物体中,但化脓性链球菌(Streptococcus pyogenes)Cas9(SpCas9)体积较大(4.3 kb),因此使用载体传递较为不便。本研究利用密码子优化后的金黄色葡萄球菌(Staphylococcus aureus)Cas9(SaCas9)系统进行了青鳉(Oryzias latipes)的tyr、oca2和pax6.1基因编辑。SaCas9片段(3.3 kb)要小,所需的原型间隔相邻基序(PAM)序列为5'-NNGRRT-3'。此外,本研究还利用tRNA-sgRNA系统在体内和体外转录表达功能性sgRNA,经SaCas9和tRNA-sgRNA的组合编辑青鳉基因组中的tyr基因。实验结果表明,SaCas9/sgRNA和SaCas9/tRNA-sgRNA系统均能有效编辑青鳉基因组,而PAM序列是该系统进行有效编辑的关键部分。此外,tRNA还能使sgRNA受巨细胞病毒等常见启动子的控制,从而提高系统的适应性。本研究还发现,CMV-SaCas9-tRNA-sgRNA-tRNA一体化结构在青鳉基因编辑中同样能发挥作用。综上,经密码子优化的SaCas9系统为编辑青鳉及潜在的其他鱼类基因组提供了一种更便捷的工具。
Keywords: Gene editing; Medaka; Oculocutaneous albinism II (oca2); Paired box 6.1 (pax6.1); Staphylococcus aureus Cas9 (SaCas9); Transfer RNA(tRNA); Tyrosinase (tyr).
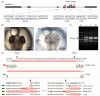
Figures







Similar articles
-
Efficient gene editing in a medaka (Oryzias latipes) cell line and embryos by SpCas9/tRNA-gRNA.J Zhejiang Univ Sci B. 2022 Jan 15;23(1):74-83. doi: 10.1631/jzus.B2100343. J Zhejiang Univ Sci B. 2022. PMID: 35029089 Free PMC article.
-
Superior Fidelity and Distinct Editing Outcomes of SaCas9 Compared with SpCas9 in Genome Editing.Genomics Proteomics Bioinformatics. 2023 Dec;21(6):1206-1220. doi: 10.1016/j.gpb.2022.12.003. Epub 2022 Dec 20. Genomics Proteomics Bioinformatics. 2023. PMID: 36549468 Free PMC article.
-
Efficient Production of Gene-Modified Mice using Staphylococcus aureus Cas9.Sci Rep. 2016 Sep 2;6:32565. doi: 10.1038/srep32565. Sci Rep. 2016. PMID: 27586692 Free PMC article.
-
Genome Editing with CRISPR-Cas9: Can It Get Any Better?J Genet Genomics. 2016 May 20;43(5):239-50. doi: 10.1016/j.jgg.2016.04.008. Epub 2016 Apr 24. J Genet Genomics. 2016. PMID: 27210042 Free PMC article. Review.
-
[CRISPR/CAS9, the King of Genome Editing Tools].Mol Biol (Mosk). 2017 Jul-Aug;51(4):582-594. doi: 10.7868/S0026898417040036. Mol Biol (Mosk). 2017. PMID: 28900076 Review. Russian.
Cited by
-
Small fish making a big difference: beloved star of environmental toxicology research in the current era.J Zhejiang Univ Sci B. 2025 Jul 15;26(7):613-632. doi: 10.1631/jzus.B2500166. J Zhejiang Univ Sci B. 2025. PMID: 40722242 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

