Host-microbe computational proteomic landscape in oral cancer revealed key functional and metabolic pathways between Fusobacterium nucleatum and cancer progression
- PMID: 39743544
- PMCID: PMC11693762
- DOI: 10.1038/s41368-024-00326-8
Host-microbe computational proteomic landscape in oral cancer revealed key functional and metabolic pathways between Fusobacterium nucleatum and cancer progression
Abstract
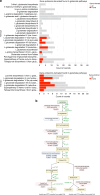
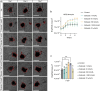
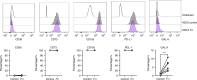
Oral squamous cell carcinoma (OSCC) is the most common manifestation of oral cancer. It has been proposed that periodontal pathogens contribute to OSCC progression, mainly by their virulence factors. However, the main periodontal pathogen and its mechanism to modulate OSCC cells remains not fully understood. In this study we investigate the main host-pathogen pathways in OSCC by computational proteomics and the mechanism behind cancer progression by the oral microbiome. The main host-pathogen pathways were analyzed in the secretome of biopsies from patients with OSCC and healthy controls by mass spectrometry. Then, functional assays were performed to evaluate the host-pathogen pathways highlighted in oral cancer. Host proteins associated with LPS response, cell migration/adhesion, and metabolism of amino acids were significantly upregulated in the human cancer proteome, whereas the complement cascade was downregulated in malignant samples. Then, the microbiome analysis revealed large number and variety of peptides from Fusobacterium nucleatum (F. nucleatum) in OSCC samples, from which several enzymes from the L-glutamate degradation pathway were found, indicating that L-glutamate from cancer cells is used as an energy source, and catabolized into butyrate by the bacteria. In fact, we observed that F. nucleatum modulates the cystine/glutamate antiporter in an OSCC cell line by increasing SLC7A11 expression, promoting L-glutamate efflux and favoring bacterial infection. Finally, our results showed that F. nucleatum and its metabolic derivates promote tumor spheroids growth, spheroids-derived cell detachment, epithelial-mesenchymal transition and Galectin-9 upregulation. Altogether, F. nucleatum promotes pro-tumoral mechanism in oral cancer.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures







References
-
- Ferlay, J. et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer144, 1941–1953 (2019). - PubMed
-
- Chow, L. Q. M. Head and Neck Cancer. N. Engl. J. Med382, 60–72 (2020). - PubMed
-
- Bremmer, J. F. et al. Prognostic value of DNA ploidy status in patients with oral leukoplakia. Oral. Oncol.47, 956–960 (2011). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

