Targeting cancer with precision: strategical insights into TCR-engineered T cell therapies
- PMID: 39744228
- PMCID: PMC11667231
- DOI: 10.7150/thno.104594
Targeting cancer with precision: strategical insights into TCR-engineered T cell therapies
Abstract
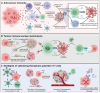
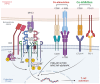
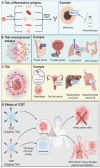
T cell receptor-engineered T (TCR-T) cell therapies are at the forefront of cancer immunotherapy, offering a transformative approach that significantly enhances the ability of T cells to recognize and eliminate cancer cells. This innovative method involves genetically modifying TCRs to increase their affinity for tumor-specific antigens. While these enhancements improve the ability of T cells to recognize and bind to antigens on cancer cells, rigorous assessment of specificity remains crucial to ensure safety and targeted responses. This dual focus on affinity and specificity holds significant promise for the treatment of solid tumors, enabling precise and efficient cancer cell recognition. Despite rapid advancements in TCR engineering and notable progress in TCR screening technologies, as evidenced by the growing number of specific TCRs entering clinical trials, several technical and clinical challenges remain. These challenges primarily pertain to the specificity, affinity, and safety of engineered TCRs. Moreover, the accurate identification and selection of TCRs that are both effective and safe are essential for the success of TCR-T cell therapies in cancer treatment. This review provides a comprehensive examination of the theoretical foundations of TCR therapy, explores strategies for screening specific TCRs and antigens, and highlights the ongoing challenges in this evolving therapeutic landscape.
Keywords: T cell receptor-engineered T cell therapy; cancer immunotherapy; screening strategy.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures



References
-
- Jhunjhunwala S, Hammer C, Delamarre L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer. 2021;21:298–312. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

