PIM1 instigates endothelial-to-mesenchymal transition to aggravate atherosclerosis
- PMID: 39744686
- PMCID: PMC11671384
- DOI: 10.7150/thno.102597
PIM1 instigates endothelial-to-mesenchymal transition to aggravate atherosclerosis
Abstract
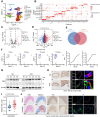
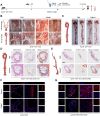
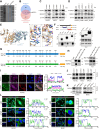
Background: Endothelial-to-mesenchymal transition (EndMT) is a cellular reprogramming mechanism by which endothelial cells acquire a mesenchymal phenotype. Endothelial cell dysfunction is the initiating factor of atherosclerosis (AS). Increasing evidence suggests that EndMT contributes to the occurrence and progression of atherosclerotic lesions and plaque instability. However, the mechanisms leading to EndMT in atherosclerotic plaques' microenvironment are poorly understood. Methods: Single-cell sequencing data of atherosclerotic plaques in mice fed with high-fat diet for different time periods were analyzed. Using quantitative polymerase chain reaction, western blotting, and immunohistochemistry, we demonstrated that the expression of PIM1 in ox-LDL stimulated endothelial cells and in human and mouse atherosclerotic lesions. ApoE -/- C57 mice were injected recombinant adeno-associated virus serotype 9 through tail vein to explore the role of PIM1 in atherosclerosis. Co-immunoprecipitation (Co-IP) was used to verify the substrates of PIM1. Hematoxylin and eosin (H&E) staining, Oil Red O staining, and Masson's trichrome staining were used to assess the size of atherosclerotic plaques, lipid content, and collagen fiber content, respectively. Results: PIM1 expression in endothelial cells increased with the progression of AS in vivo. Endothelial cell-specific PIM1 knockdown negatively regulated atherosclerosis progression and the EndMT process. Knockdown of PIM1 in endothelial cells in vitro attenuated ox-LDL-induced EndMT. This process was primarily due to the reduction of PIM1, which led to decreased phosphorylation of NDRG1 at Ser330, and subsequently, reduced NDRG1 nuclear translocation. Consequently, the interaction between NDRG1 and PTBP1 was affected, ultimately impacting the mRNA levels of Vimentin, ZEB1, Slug, Snail, N-Cadherin, TAGLN, and α-SMA. The small molecule Max-40279 could inhibit NDRG1 phosphorylation at Ser330 and suppress EndMT. Conclusion: Our findings revealed the PIM1/P-NDRG1(S330)/PTBP1/EndMT axis as a critical factor promoting AS progression and could generate new strategies to prevent AS.
Keywords: Atherosclerosis; Endothelial-to-mesenchymal transition; Max-40279; Nuclear translocation; PIM1.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures





References
-
- Li A, Peng W, Xia X, Li R, Wang Y, Wei D. Endothelial-to-Mesenchymal Transition: A Potential Mechanism for Atherosclerosis Plaque Progression and Destabilization. DNA Cell Biol. 2017;36:883–91. - PubMed
-
- Madar S, Goldstein I, Rotter V. 'Cancer associated fibroblasts'-more than meets the eye. Trends Mol Med. 2013;19:447–53. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

