Single-cell data reveal heterogeneity of investment in ribosomes across a bacterial population
- PMID: 39746998
- PMCID: PMC11695989
- DOI: 10.1038/s41467-024-55394-5
Single-cell data reveal heterogeneity of investment in ribosomes across a bacterial population
Abstract
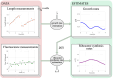
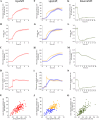
Ribosomes are responsible for the synthesis of proteins, the major component of cellular biomass. Classical experiments have established a linear relationship between the fraction of resources invested in ribosomal proteins and the rate of balanced growth of a microbial population. Very little is known, however, about how the investment in ribosomes varies over individual cells in a population. We therefore extended the study of ribosomal resource allocation from populations to single cells, using a combination of time-lapse fluorescence microscopy and statistical inference. We found a large variability of ribosome concentrations and growth rates in conditions of balanced growth of the model bacterium Escherichia coli in a given medium, which cannot be accounted for by the population-level growth law. A large variability in the allocation of resources to ribosomes was also found during the transition of the bacteria from a poor to a rich growth medium. While some cells immediately adapt their ribosome synthesis rate to the new environment, others do so only gradually. Our results thus reveal a range of strategies for investing resources in the molecular machines at the heart of cellular self-replication. This raises the fundamental question whether the observed variability is an intrinsic consequence of the stochastic nature of the underlying biochemical processes or whether it improves the fitness of Escherichia coli in its natural environment.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



References
-
- Schaechter, M., Maaløe, O. & Kjeldgaard, O. N. Dependency on medium and temperature of cell size and chemical composition during balanced growth of Salmonella typhimurium. J. Gen. Microbiol. 19, 592–606 (1958). - PubMed
-
- Neidhardt, F. C. & Magasanik, B. Studies on the role of ribonucleic acid in the growth of bacteria. Biochim. Biophys. Acta42, 99–116 (1960). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

