Parameters for one health genomic surveillance of Escherichia coli from Australia
- PMID: 39747833
- PMCID: PMC11696363
- DOI: 10.1038/s41467-024-55103-2
Parameters for one health genomic surveillance of Escherichia coli from Australia
Erratum in
-
Publisher Correction: Parameters for one health genomic surveillance of Escherichia coli from Australia.Nat Commun. 2025 Apr 7;16(1):3309. doi: 10.1038/s41467-025-58600-0. Nat Commun. 2025. PMID: 40195308 Free PMC article. No abstract available.
Abstract
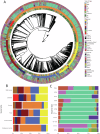
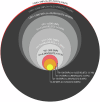
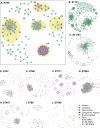
Genomics is a cornerstone of modern pathogen epidemiology yet demonstrating transmission in a One Health context is challenging, as strains circulate and evolve within and between diverse hosts and environments. To identify phylogenetic linkages and better define relevant measures of genomic relatedness in a One Health context, we collated 5471 Escherichia coli genome sequences from Australia originating from humans (n = 2996), wild animals (n = 870), livestock (n = 649), companion animals (n = 375), environmental sources (n = 292) and food (n = 289) spanning over 36 years. Of the 827 multi-locus sequence types (STs) identified, 10 STs were commonly associated with cross-source genomic clusters, including the highly clonal ST131, pandemic zoonotic lineages such as ST95, and emerging human ExPEC ST1193. Here, we show that assessing genomic relationships at ≤ 100 SNP threshold enabled detection of cross-source linkage otherwise obscured when applying typical outbreak-oriented relatedness thresholds ( ≤ 20 SNPs) and should be considered in interrogation of One Health genomic datasets.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare that there are no conflicts of interest regarding the generation and publication of this manuscript.
Figures


References
-
- Nagy, B. & Fekete, P. Z. Enterotoxigenic Escherichia coli (ETEC) in farm animals. Vet. Res30, 259–284 (1999). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

