FN1 shapes the behavior of papillary thyroid carcinoma through alternative splicing of EDB region
- PMID: 39747903
- PMCID: PMC11695688
- DOI: 10.1038/s41598-024-83369-5
FN1 shapes the behavior of papillary thyroid carcinoma through alternative splicing of EDB region
Abstract
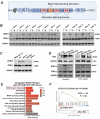
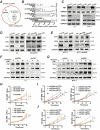
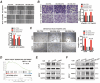
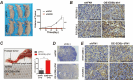
Papillary thyroid cancer (PTC) is often characterized by indolent behavior, small tumors with slow cell proliferation and a tendency to metastasize to cervical lymph node simultaneously, and the molecular mechanisms underlying that remain poorly understood. In this study, FN1 was the hottest gene of PTC and distinctive expression in PTC cells. FN1 deficiency severely inhibited the p53 signaling pathway, especially cyclin proteins, resulting in increased cell growth but hampered invasion. The alternatively splicing EDB region of FN1 was exclusively expressed in tumors, which impacted integrin β1 (ITGB1) bonding FN1 and its secretion process, resulting in completely distinct roles of two isoforms that FN1 including and skipping EDB domain. The isoform EDB(-)FN1 intracellularly inhibited tumor proliferation by upregulating p21 expression, whereas extracellular EDB(+)FN1 promoted lymph node metastasis via the VEGF signaling pathway in vitro and in vivo. Moreover, the alternative splicing EDB region of FN1 was modulated by p53-targeted protein ZMAT3 which activated cell migration and lymphoangiogenesis. Collectively, combined with p53-induced proteins, FN1 played both anti- and pro-cancer roles owing to EDB domain alternative splicing. FN1 is a potential determinant behind the characteristic behavior of PTC, which may contribute to a deeper understanding of the peculiarity of PTC and provide a promising target for regional lymph node metastasis.
Keywords: Alternative splicing EDB domain; FN1; Lymph node metastasis; Papillary thyroid cancer; ZMAT3; p53/p21.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Ethical approval was provided by the Committees for Ethical Review of Research involving Human Subjects and Animal experiments at Xiangya Hospital, Central South University (No. 2019030140). Competing interests: The authors declare no competing interests.
Figures




Similar articles
-
Integrin-fibronectin interaction is a pivotal biological and clinical determinant in papillary thyroid carcinoma.Endocr Relat Cancer. 2025 Jun 5;32(6):e250101. doi: 10.1530/ERC-25-0101. Print 2025 Jun 1. Endocr Relat Cancer. 2025. PMID: 40423510 Free PMC article.
-
MicroRNA-142-3P suppresses the progression of papillary thyroid carcinoma by targeting FN1 and inactivating FAK/ERK/PI3K signaling.Cell Signal. 2023 Sep;109:110792. doi: 10.1016/j.cellsig.2023.110792. Epub 2023 Jul 3. Cell Signal. 2023. PMID: 37406787
-
Functional Characterization of Plasminogen Activator Urokinase as a Key Gene in Papillary Thyroid Carcinoma Lymph Node Metastasis.Genet Test Mol Biomarkers. 2025 Jun;29(6):152-165. doi: 10.1089/gtmb.2025.0046. Epub 2025 Jun 2. Genet Test Mol Biomarkers. 2025. PMID: 40454923
-
FBXO2 promotes the progression of papillary thyroid carcinoma through the p53 pathway.Sci Rep. 2024 Sep 29;14(1):22574. doi: 10.1038/s41598-024-73455-z. Sci Rep. 2024. PMID: 39343799 Free PMC article.
-
Fibronectin-targeted drug delivery in cancer.Adv Drug Deliv Rev. 2016 Feb 1;97:101-10. doi: 10.1016/j.addr.2015.11.014. Epub 2015 Nov 27. Adv Drug Deliv Rev. 2016. PMID: 26639577 Review.
References
-
- Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. Ca-a Cancer J. Clin.72 (1), 7–33 (2022). - PubMed
-
- Miranda, A. et al. Thyroid cancer incidence trends by histology in 25 countries: a population-based study. Lancet Diabetes Endocrinol.9 (4), 225–234 (2021). - PubMed
-
- Schlumberger, M. & Leboulleux, S. Current practice in patients with differentiated thyroid cancer. Nat. Reviews Endocrinol.17 (3), 176–188 (2021). - PubMed
-
- Yu, X., Wan, Y., Sippel, R. & Chen, H. J. A. o. s., Should all papillary thyroid microcarcinomas be aggressively treated? An analysis of 18,445 cases. Ann. Surg.254 (4), 653–660 (2011). - PubMed
-
- Sciuto, R. et al. Natural history and clinical outcome of differentiated thyroid carcinoma: a retrospective analysis of 1503 patients treated at a single institution. Ann. Oncol.20 (10), 1728–1735 (2009). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

