Prediction of Patient Drug Response via 3D Bioprinted Gastric Cancer Model Utilized Patient-Derived Tissue Laden Tissue-Specific Bioink
- PMID: 39748450
- PMCID: PMC11905052
- DOI: 10.1002/advs.202411769
Prediction of Patient Drug Response via 3D Bioprinted Gastric Cancer Model Utilized Patient-Derived Tissue Laden Tissue-Specific Bioink
Abstract
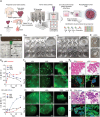
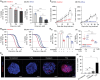
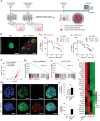
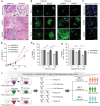
Despite significant research progress, tumor heterogeneity remains elusive, and its complexity poses a barrier to anticancer drug discovery and cancer treatment. Response to the same drug varies across patients, and the timing of treatment is an important factor in determining prognosis. Therefore, development of patient-specific preclinical models that can predict a patient's drug response within a short period is imperative. In this study, a printed gastric cancer (pGC) model is developed for preclinical chemotherapy using extrusion-based 3D bioprinting technology and tissue-specific bioinks containing patient-derived tumor chunks. The pGC model retained the original tumor characteristics and enabled rapid drug evaluation within 2 weeks of its isolation from the patient. In fact, it is confirmed that the drug response-related gene profile of pGC tissues co-cultured with human gastric fibroblasts (hGaFibro) is similar to that of patient tissues. This suggested that the application of the pGC model can potentially overcome the challenges associated with accurate drug evaluation in preclinical models (e.g., patient-derived xenografts) owing to the deficiency of stromal cells derived from the patient. Consequently, the pGC model manifested a remarkable similarity with patients in terms of response to chemotherapy and prognostic predictability. Hence, it is considered a promising preclinical tool for personalized and precise treatments.
Keywords: drug efficacy testing; gastric cancer patient‐derived xenograft; gastric tissue‐derived decellularized extracellular matrix; tumor tissue printing.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Gastric Cancer Models Developed via GelMA 3D Bioprinting Accurately Mimic Cancer Hallmarks, Tumor Microenvironment Features, and Drug Responses.Small. 2025 Feb;21(8):e2409321. doi: 10.1002/smll.202409321. Epub 2025 Jan 15. Small. 2025. PMID: 39811968
-
Alginate-Based Bioinks for 3D Bioprinting and Fabrication of Anatomically Accurate Bone Grafts.Tissue Eng Part A. 2021 Sep;27(17-18):1168-1181. doi: 10.1089/ten.TEA.2020.0305. Epub 2021 Feb 26. Tissue Eng Part A. 2021. PMID: 33218292 Free PMC article.
-
Bioprinting 101: Design, Fabrication, and Evaluation of Cell-Laden 3D Bioprinted Scaffolds.Tissue Eng Part A. 2020 Mar;26(5-6):318-338. doi: 10.1089/ten.TEA.2019.0298. Tissue Eng Part A. 2020. PMID: 32079490 Free PMC article.
-
Three-Dimensional Bioprinting of Decellularized Extracellular Matrix-Based Bioinks for Tissue Engineering.Molecules. 2022 May 26;27(11):3442. doi: 10.3390/molecules27113442. Molecules. 2022. PMID: 35684380 Free PMC article. Review.
-
Bioink: a 3D-bioprinting tool for anticancer drug discovery and cancer management.Drug Discov Today. 2021 Jul;26(7):1574-1590. doi: 10.1016/j.drudis.2021.03.010. Epub 2021 Mar 16. Drug Discov Today. 2021. PMID: 33741496 Review.
Cited by
-
Biomaterial-driven regenerative drug delivery: a vicennial bibliometric landscape.Front Med (Lausanne). 2025 Jul 14;12:1593985. doi: 10.3389/fmed.2025.1593985. eCollection 2025. Front Med (Lausanne). 2025. PMID: 40727534 Free PMC article.
-
Three-Dimensional Culture System: A New Frontier in Cancer Research, Drug Discovery, and Stem Cell-Based Therapy.Biology (Basel). 2025 Jul 17;14(7):875. doi: 10.3390/biology14070875. Biology (Basel). 2025. PMID: 40723433 Free PMC article. Review.
References
-
- Pleasance E., Bohm A., Williamson L. M., Nelson J. M. T., Shen Y., Bonakdar M., Titmuss E., Csizmok V., Wee K., Hosseinzadeh S., Grisdale C. J., Reisle C., Taylor G. A., Lewis E., Jones M. R., Bleile D., Sadeghi S., Zhang W., Davies A., Pellegrini B., Wong T., Bowlby R., Chan S. K., Mungall K. L., Chuah E., Mungall A. J., Moore R. A., Zhao Y., Deol B., Fisic A., et al., Ann. Oncol. 2022, 33, 939. - PubMed
-
- Patch A.‐M., Christie E. L., Etemadmoghadam D., Garsed D. W., George J., Fereday S., Nones K., Cowin P., Alsop K., Bailey P. J., Kassahn K. S., Newell F., Quinn M. C. J., Kazakoff S., Quek K., Wilhelm‐Benartzi C., Curry E., Leong H. S., Hamilton A., Mileshkin L., Au‐Yeung G., Kennedy C., Hung J., Chiew Y.‐E., Harnett P., Friedlander M., Quinn M., Pyman J., Cordner S., O'Brien P., et al., Nature 2015, 521, 489. - PubMed
MeSH terms
Substances
Grants and funding
- 2020R1A6A1A03047902/Ministry of Education, Basic Science Research Program through the National Research Foundation of Korea(NRF)
- 2022M3C1A3081359/Korea government (MSIT), National Research Foundation of Korea(NRF) grant.
- 2021R1A2C2004981/Korea government (MSIT), National Research Foundation of Korea(NRF) grant.
LinkOut - more resources
Full Text Sources
Medical
