Deeper response predicts better outcomes in high-risk-smoldering-myeloma: results of the I-PRISM phase II clinical trial
- PMID: 39753553
- PMCID: PMC11698957
- DOI: 10.1038/s41467-024-55308-5
Deeper response predicts better outcomes in high-risk-smoldering-myeloma: results of the I-PRISM phase II clinical trial
Abstract
Early therapeutic intervention in high-risk smoldering multiple myeloma (HR-SMM) has shown benefits, however, no studies have assessed whether biochemical progression or response depth predicts long-term outcomes. The single-arm I-PRISM phase II trial (NCT02916771) evaluated ixazomib, lenalidomide, and dexamethasone in 55 patients with HR-SMM. The primary endpoint, median progression-free survival (PFS), was not reached (NR) (95% CI: 57.7-NR, median follow-up 50 months). The secondary endpoint, biochemical PFS, was 48.6 months (95% CI: 39.9-NR) and coincided with or preceded SLiM-CRAB in eight patients. For additional secondary objectives, the overall response rate was 93% with 31% achieving complete response (CR) and 45% very good partial response (VGPR) or better. CR correlated strongly with the absence of SLiM-CRAB and biochemical progression. MRD-negativity (10-5 sensitivity) predicted a 5-year biochemical PFS of 100% versus 40% in MRD-positive patients (p = 0.051), demonstrating that deep responses significantly improve time to progression. Exploratory single-cell RNA sequencing linked tumor MHC class I expression to proteasome inhibitor response, and a lower proportion of GZMB+ T cells within clonally expanded CD8+ T cells associated with suboptimal outcomes.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: O.N. reports research support from Takeda and Janssen; advisory board participation for Bristol Myers Squibb, Janssen, Sanofi, Takeda, and GPCR therapeutics; and honorarium from Pfizer. M.B. reports consultancy for Janssen, BMS, Takeda, Epizyme, Karyopharm, Menarini Biosystems, and Adaptive. G.B. reports consultancy for Prothena. E.O. reports advisory board participation and/or honoraria from Janssen, BMS, Sanofi, Pfizer, Exact; consulting for Takeda; and steering committee participation with Natera. K.A. reports consultancy for AstraZeneca, Janssen, and Pfizer; and board participation and stock options for Dynamic Cell Therapies, C4 Therapeutics, Next RNA, Oncopep, Starton, and Window. P.G.R. reports advisory board participation or consulting for Celgene (BMS), GSK, Karyopharm, Oncopeptides, Regeneron, Sanofi, and Takeda; and research grants from Oncopeptides and Karyopharm. R.S.P. is a co-founder, equity holder, and consultant for PreDICTA Bioscience. I.M.G.: Consulting/Advisory role: AbbVie, Adaptive, Amgen, Aptitude Health, Bristol Myers Squibb, GlaxoSmithKline, Huron Consulting, Janssen, Menarini Silicon Biosystems, Oncopeptides, Pfizer, Sanofi, Sognef, Takeda, Binding Site a part of Thermo Fischer Scientific, Window Therapeutics, and 10X Genomics; speaker fees from Vor Biopharma and Veeva Systems, Inc., and is a co-founder of PreDICTA Bioscience. I.M.G.’s spouse is the CMO and an equity holder of Disc Medicine. All other authors declare no conflicts of interest. Inclusion and ethics: One or more of the authors of this paper self-identifies as an underrepresented ethnic and/or gender minority in science. One or more of the authors of this paper self-identifies as a member of the LGBTQIA+ community. We support inclusive, diverse, and equitable conduct of research.
Figures



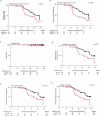


References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

