Expanding the immune-related targetome of miR-155-5p by integrating time-resolved RNA patterns into miRNA target prediction
- PMID: 39760255
- PMCID: PMC11730359
- DOI: 10.1080/15476286.2025.2449775
Expanding the immune-related targetome of miR-155-5p by integrating time-resolved RNA patterns into miRNA target prediction
Abstract
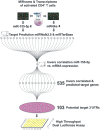
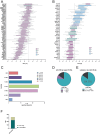
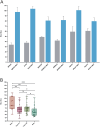
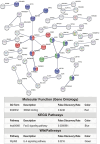
The lack of a sufficient number of validated miRNA targets severely hampers the understanding of their biological function. Even for the well-studied miR-155-5p, there are only 239 experimentally validated targets out of 42,554 predicted targets. For a more complete assessment of the immune-related miR-155 targetome, we used an inverse correlation of time-resolved mRNA profiles and miR-155-5p expression of early CD4+ T cell activation to predict immune-related target genes. Using a high-throughput miRNA interaction reporter (HiTmIR) assay we examined 90 target genes and confirmed 80 genes as direct targets of miR-155-5p. Our study increases the current number of verified miR-155-5p targets approximately threefold and exemplifies a method for verifying miRNA targetomes as a prerequisite for the analysis of miRNA-regulated cellular networks.
Keywords: CD4+ T cell activation; miR-155-5p; miRnomics; target prediction; time-resolved RNA patterns; transcriptomics.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
