Unravelling distinct patterns of metagenomic surveillance and respiratory microbiota between two P1 genotypes of Mycoplasma pneumoniae
- PMID: 39760260
- PMCID: PMC11730683
- DOI: 10.1080/22221751.2024.2449087
Unravelling distinct patterns of metagenomic surveillance and respiratory microbiota between two P1 genotypes of Mycoplasma pneumoniae
Abstract
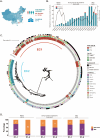
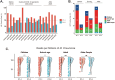
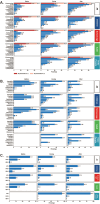
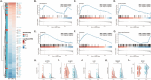
To unravel distinct patterns of metagenomic surveillance and respiratory microbiota between Mycoplasma pneumoniae (M. pneumoniae) P1-1 and P1-2 and to explore the impact of the COVID-19 pandemic on epidemiological features, we conducted a multicentre retrospective study which spanned 90,886 pneumonia patients, among which 3164 cases M. pneumoniae were identified. Our findings revealed a concurrent outbreak of M. pneumoniae, with the positivity rate rising sharply to 9.62% from July 2023, compared to the 0.16% to 4.06% positivity rate observed during the 2020-2022 COVID-19 pandemic. P1-1 had a higher odds ratio of co-detecting opportunistic pathogens. However, no significant differences were observed in the co-detection odds ratio between children and other age groups in P1-2. This study is the first to demonstrate differences in relative abundance, diversity of respiratory microbiota and co-detection rate of opportunistic pathogen between M. pneumoniae P1-1 and P1-2. Through bronchoalveolar lavage (BAL) metagenomic and host transcriptomic analyses, we identified variations in co-detection rates of M. pneumoniae P1-1 genotype with opportunistic pathogens like S. pneumoniae, alterations in respiratory microbiota composition, lung inflammation, and disruption of ciliary function. Consistent with the results of host transcriptome, we found that P1-1 infections were associated with significantly higher rates of requiring respiratory support and mechanical ventilation compared to P1-2 infections (Fisher's exact test, p-value = 0.035/0.004). Our study provides preliminary evidence of clinical severity between M. pneumoniae strains, underscoring the need for ongoing research and development of targeted therapeutic strategies.
Keywords: Mycoplasma pneumoniae; P1 genotype; clinical outcome; host immune response; respiratory microbiota.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
