Discovery of a parallel family of euglenatide analogs in Euglena gracilis
- PMID: 39760919
- PMCID: PMC11703798
- DOI: 10.1007/s13659-024-00490-8
Discovery of a parallel family of euglenatide analogs in Euglena gracilis
Abstract
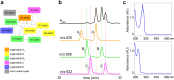
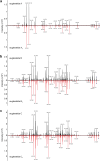
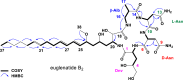
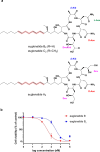
The euglenatides are a family of hybrid polyketide-nonribosomal peptides produced by the unicellular algae Euglena gracilis. These compounds have antiproliferative activity against fungal pathogens and mammalian cancer cell lines. Analysis of E. gracilis extracts revealed that the algae produce not only the euglenatides, but also a corresponding family of analogs that have the same molecular weights as the euglenatides, but are lacking the characteristic triene chromophore. In comparison to the euglenatides, the activity of these analogs is greatly reduced in a mammalian cytotoxicity assay, indicating that the triene is critical to the biological activity of the euglenatides.
Keywords: Euglena gracilis; Euglenatide; Natural products; Nonribosomal peptide; Polyketide.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no conflict of interest.
Figures

Similar articles
-
Euglenatides, Potent Antiproliferative Cyclic Peptides Isolated from the Freshwater Photosynthetic Microalga Euglena gracilis.Angew Chem Int Ed Engl. 2022 Jun 7;61(23):e202203175. doi: 10.1002/anie.202203175. Epub 2022 Apr 6. Angew Chem Int Ed Engl. 2022. PMID: 35325497 Free PMC article.
-
Gravity sedimentation of eukaryotic algae Euglena gracilis accelerated by ethanol cultivation.Appl Microbiol Biotechnol. 2023 May;107(9):3021-3032. doi: 10.1007/s00253-023-12476-6. Epub 2023 Mar 21. Appl Microbiol Biotechnol. 2023. PMID: 36941437
-
Fast and efficient molecule delivery into Euglena gracilis mediated by cell-penetrating peptide or dimethyl sulfoxide.FEBS Open Bio. 2023 Apr;13(4):597-605. doi: 10.1002/2211-5463.13592. Epub 2023 Mar 18. FEBS Open Bio. 2023. PMID: 36883745 Free PMC article.
-
[Advances of studies on culture and product functions of Euglena gracilis].Sheng Wu Gong Cheng Xue Bao. 2024 Mar 25;40(3):705-721. doi: 10.13345/j.cjb.230309. Sheng Wu Gong Cheng Xue Bao. 2024. PMID: 38545972 Review. Chinese.
-
The biomolecules of Euglena gracilis: Harnessing biology for natural solutions to future problems.Protist. 2024 Aug;175(4):126044. doi: 10.1016/j.protis.2024.126044. Epub 2024 May 23. Protist. 2024. PMID: 38823247 Review.
References
-
- Miyanaga A, Kudo F, Eguchi T. Protein-protein interactions in polyketide synthase-nonribosomal peptide synthetase hybrid assembly lines. Nat Prod Rep. 2018;35:1185–209. 10.1039/c8np00022k. - PubMed
-
- O’Neill EC, Trick M, Hill L, Rejzek M, Dusi RG, Hamilton CJ, Zimba PV, Henrissat B, Field RA. The transcriptome of Euglena gracilis reveals unexpected metabolic capabilities for carbohydrate and natural product biochemistry. Mol Biosyst. 2015;11:2808–20. 10.1039/c5mb00319a. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

