Assembly and comparative analysis of the complete mitogenome of Rubus chingii var. suavissimus, an exceptional berry plant possessing sweet leaves
- PMID: 39764230
- PMCID: PMC11701164
- DOI: 10.3389/fpls.2024.1504687
Assembly and comparative analysis of the complete mitogenome of Rubus chingii var. suavissimus, an exceptional berry plant possessing sweet leaves
Abstract
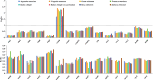
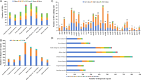
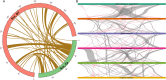
Rubus chingii var. suavissimus is a special berry plant of Rubus in the Rosaceae family. Its leaves contain high-sweetness, low-calorie, and non-toxic sweet ingredients, known as rubusoside. As a medicine and food biofunctional plant, it is a combination of "tea, sugar, and medicine." In this study, the complete mitogenome of R. chingii var. suavissimus was successfully assembled and annotated based on PacBio HiFi sequencing technology. The mitogenome of R. chingii var. suavissimus was a typical master circle structure, spanning 432,483 bp and containing 34 unique protein-coding genes (PCGs), 20 tRNAs, and 3 rRNAs. The majority of these PCGs was subjected to purifying selection, and only one gene (ccmB) showed sign of positive selection. The mitogenome of R. chingii var. suavissimus contained a large number of repeats, and the homogeneous fragments transferring between plastid genome and mitogenome, with a total of 55 pairs of mitochondrial plastid sequences (MTPTs), and the total size was 56,913 bp. Comparative analysis showed that the non-coding region in the mitogenome of R. chingii var. suavissimus had undergone frequent rearrangements during evolution, but the coding region was still highly conserved. Furthermore, the maximum likelihood and Bayesian inference phylogenetic trees were reconstructed of 10 shared PCGs in 36 plant species. The topological structures of two phylogenetic trees were consistent with the APG IV classification system and had high support rates. In general, this study clarifies the mitogenome of R. chingii var. suavissimus and provides valuable insights into the genetic evolution of the Rosaceae family.
Keywords: Rubus chingii var. suavissimus; master circle structure; mitochondrial genome; phylogenetic analysis; rearrangement.
Copyright © 2024 Shi, Chen, Jiang, Wu, Xin and Zeng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures




Similar articles
-
[Comparative analysis of plastomes of Rubus chingii and R. chingii var. suavissimus].Zhongguo Zhong Yao Za Zhi. 2024 Mar;49(6):1506-1516. doi: 10.19540/j.cnki.cjcmm.20231115.105. Zhongguo Zhong Yao Za Zhi. 2024. PMID: 38621934 Chinese.
-
Comparative Metabolome and Transcriptome Analysis Revealed the Accumulative Mechanism of Rubusoside in Chinese Sweet Tea.J Agric Food Chem. 2024 Nov 6;72(44):24539-24551. doi: 10.1021/acs.jafc.4c07127. Epub 2024 Oct 23. J Agric Food Chem. 2024. PMID: 39442010
-
Comprehensive Analysis of the Complete Mitochondrial Genome of Rehmannia chingii: An Autotrophic Species in the Orobanchaceae Family.Genes (Basel). 2024 Jan 15;15(1):98. doi: 10.3390/genes15010098. Genes (Basel). 2024. PMID: 38254987 Free PMC article.
-
Bioactive components, pharmacological effects, and drug development of traditional herbal medicine Rubus chingii Hu (Fu-Pen-Zi).Front Nutr. 2023 Jan 9;9:1052504. doi: 10.3389/fnut.2022.1052504. eCollection 2022. Front Nutr. 2023. PMID: 36698464 Free PMC article. Review.
-
Rubus chingii Hu: an overview of botany, traditional uses, phytochemistry, and pharmacology.Chin J Nat Med. 2020 Jun;18(6):401-416. doi: 10.1016/S1875-5364(20)30048-0. Chin J Nat Med. 2020. PMID: 32503732
Cited by
-
Complete sequencing of the mitochondrial genome of tea plant Camellia sinensis cv. 'Baihaozao': multichromosomal structure, phylogenetic relationships, and adaptive evolutionary analysis.Front Plant Sci. 2025 Jun 13;16:1604404. doi: 10.3389/fpls.2025.1604404. eCollection 2025. Front Plant Sci. 2025. PMID: 40584852 Free PMC article.
-
Complete plastome and mitogenome assembly of endangered tree Karpatiosorbus bristoliensis reveals phylogenetic architecture for Sorbus sensu lato (Rosaceae).Front Plant Sci. 2025 Jul 8;16:1619267. doi: 10.3389/fpls.2025.1619267. eCollection 2025. Front Plant Sci. 2025. PMID: 40697870 Free PMC article.
-
Mitogenome Characteristics and Intracellular Gene Transfer Analysis of Four Adansonia Species.Genes (Basel). 2025 Jul 21;16(7):846. doi: 10.3390/genes16070846. Genes (Basel). 2025. PMID: 40725502 Free PMC article.
-
De-novo assembly and comparative analysis of the complete mitogenome of traditional Chinese medicine Strobilanthes sarcorrhiza.BMC Plant Biol. 2025 May 21;25(1):675. doi: 10.1186/s12870-025-06731-3. BMC Plant Biol. 2025. PMID: 40399802 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Research Materials

