This is a preprint.
The IBEX Imaging Knowledge-Base: A Community Resource Enabling Adoption and Development of Immunofluoresence Imaging Methods
- PMID: 39764400
- PMCID: PMC11702815
The IBEX Imaging Knowledge-Base: A Community Resource Enabling Adoption and Development of Immunofluoresence Imaging Methods
Abstract
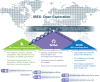
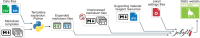
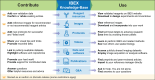
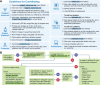
The iterative bleaching extends multiplexity (IBEX) Knowledge-Base is a central portal for researchers adopting IBEX and related 2D and 3D immunofluorescence imaging methods. The design of the Knowledge-Base is modeled after efforts in the open-source software community and includes three facets: a development platform (GitHub), static website, and service for data archiving. The Knowledge-Base facilitates the practice of open science throughout the research life cycle by providing validation data for recommended and non-recommended reagents, e.g., primary and secondary antibodies. In addition to reporting negative data, the Knowledge-Base empowers method adoption and evolution by providing a venue for sharing protocols, videos, datasets, software, and publications. A dedicated discussion forum fosters a sense of community among researchers while addressing questions not covered in published manuscripts. Together, scientists from around the world are advancing scientific discovery at a faster pace, reducing wasted time and effort, and instilling greater confidence in the resulting data.
Figures
References
-
- Bandrowski A, Brush M, Grethe JS, Haendel MA, Kennedy DN, Hill S, Hof PR, Martone ME, Pols M, Tan SC, Washington N, Zudilova-Seinstra E, Vasilevsky N. The Resource Identification Initiative: A cultural shift in publishing. Journal of Comparative Neurology. 2016; 524(1):8–22. doi: 10.1002/brb3.417. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
