Analyses of Human Genetic Data to Identify Clinically Relevant Domains of Neuroligins
- PMID: 39766868
- PMCID: PMC11675371
- DOI: 10.3390/genes15121601
Analyses of Human Genetic Data to Identify Clinically Relevant Domains of Neuroligins
Abstract
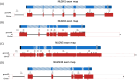
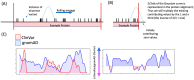
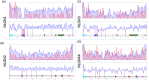
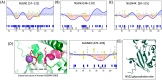
Background/Objectives: Neuroligins (NLGNs) are postsynaptic adhesion molecules critical for neuronal development that are highly associated with autism spectrum disorder (ASD). Here, we provide an overview of the literature on NLGN rare variants. In addition, we introduce a new approach to analyze human variation within NLGN genes to identify sensitive regions that have an increased frequency of ASD-associated variants to better understand NLGN function. Methods: To identify critical protein subdomains within the NLGN gene family, we developed an algorithm that assesses tolerance to missense mutations in human genetic variation by comparing clinical variants from ClinVar to reference variants from gnomAD. This approach provides tolerance values to subdomains within the protein. Results: Our algorithm identified several critical regions that were conserved across multiple NLGN isoforms. Importantly, this approach also identified a previously reported cluster of pathogenic variants in NLGN4X (also conserved in NLGN1 and NLGN3) as well as a region around the highly characterized NLGN3 R451C ASD-associated mutation. Additionally, we highlighted other, as of yet, uncharacterized regions enriched with mutations. Conclusions: The systematic analysis of NLGN ASD-associated variants compared to variants identified in the unaffected population (gnomAD) reveals conserved domains in NLGN isoforms that are tolerant to variation or are enriched in clinically relevant variants. Examination of databases also allows for predictions of the presumed tolerance to loss of an allele. The use of the algorithm we developed effectively allowed the evaluation of subdomains of NLGNs and can be used to examine other ASD-associated genes.
Keywords: autism spectrum disorder; haploinsufficiency; human genetic variation; missense variation; neurodevelopmental disorder; neuroligin.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
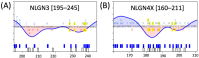
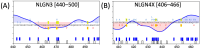
References
-
- Kenny E.M., Cormican P., Furlong S., Heron E., Kenny G., Fahey C., Kelleher E., Ennis S., Tropea D., Anney R., et al. Excess of Rare Novel Loss-of-Function Variants in Synaptic Genes in Schizophrenia and Autism Spectrum Disorders. Mol. Psychiatry. 2014;19:872–879. doi: 10.1038/mp.2013.127. - DOI - PubMed
-
- Gudmundsson S., Singer-Berk M., Watts N.A., Phu W., Goodrich J.K., Solomonson M., Genome Aggregation Database Consortium. Rehm H.L., MacArthur D.G., O’Donnell-Luria A. Variant Interpretation Using Population Databases: Lessons from gnomAD. Hum. Mutat. 2022;43:1012–1030. doi: 10.1002/humu.24309. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

