Genome-Wide Association Analysis of Growth Traits in Hu Sheep
- PMID: 39766904
- PMCID: PMC11675594
- DOI: 10.3390/genes15121637
Genome-Wide Association Analysis of Growth Traits in Hu Sheep
Abstract
(1) Background: The Hu sheep is a renowned breed characterized by high reproduction, year-round estrus, and resistance to high humidity and temperature conditions. However, the breed exhibits lower growth rates and meat yields, which necessitate improvements through selective breeding. The integration of molecular markers in sheep breeding programs has the potential to enhance growth performance, reduce breeding cycles, and increase meat production. Currently, the applications of SNP chips for genotyping in conjunction with genome-wide association studies (GWAS) have become a prevalent approach for identifying candidate genes associated with economically significant traits in livestock. (2) Methods: To pinpoint candidate genes influencing growth traits in Hu sheep, we recorded the birth weight, weaning weight, and weights at 3, 4, 5, 6, and 7 months for a total of 567 Hu sheep, and genotyping was performed using the Ovine 40K SNP chip. (3) Results: Through GWAS analysis and KEGG pathway enrichment, we identified three candidate genes associated with birth weight (CAMK2B, CACNA2D1, and CACNA1C). Additionally, we found two candidate genes linked to weaning weight (FGF9 and BMPR1B), with CACNA2D1 also serving as a shared gene between birth weight and weaning weight traits. Furthermore, we identified eight candidate genes related to monthly weight (FIGF, WT1, KCNIP4, JAK2, WWP1, PLCL1, GPRIN3, and CCSER1). (4) Conclusion: Our findings revealed a total of 13 candidate genetic markers that can be utilized for molecular marker-assisted selection, aiming to improve meat production in sheep breeding programs.
Keywords: GWAS; Hu sheep; Ovine 40K SNP chip; candidate genes; growth.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
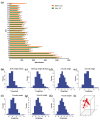


References
-
- Li X., Yang J., Shen M., Xie X.L., Liu G.J., Xu Y.X., Lv F.H., Yang H., Yang Y.L., Liu C.B., et al. Whole-Genome Resequencing of Wild and Domestic Sheep Identifies Genes Associated with Morphological and Agronomic Traits. Nat. Commun. 2020;11:2815. doi: 10.1038/s41467-020-16485-1. - DOI - PMC - PubMed
-
- Knapik J., Ropka-Molik K., Pieszka M. Genetic and Nutritional Factors Determining the Production and Quality of Sheep Meat—A Review. Ann. Anim. Sci. 2017;17:23–40. doi: 10.1515/aoas-2016-0036. - DOI
-
- Wang X., Wang Y.C., Wang Q.Y., Dai C.P., Li J.Z., Huang P.F., Li Y.L., Ding X.Q., Huang J., Hussain T., et al. The Impact of Early and Mid-Pregnant Hu Ewes’ Dietary Protein and Energy Levels on Growth Performance and Serum Biochemical Indices. J. Appl. Anim. Res. 2023;51:174–181. doi: 10.1080/09712119.2023.2170385. - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

