Genetic Diversity and Fingerprinting of 231 Mango Germplasm Using Genome SSR Markers
- PMID: 39769387
- PMCID: PMC11728225
- DOI: 10.3390/ijms252413625
Genetic Diversity and Fingerprinting of 231 Mango Germplasm Using Genome SSR Markers
Abstract
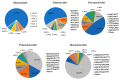
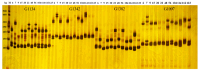
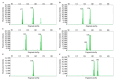
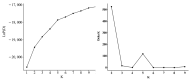
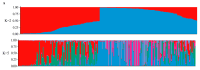
Mango (Mangifera indica L.) (2n = 40) is an important perennial fruit tree in tropical and subtropical regions. The lack of information on genetic diversity at the molecular level hinders efforts in mango genetic improvement and molecular marker-assisted breeding. In this study, a genome-wide screening was conducted to develop simple sequence repeat (SSR) markers using the Alphonso reference genome. A total of 187 SSR primer pairs were designed based on SSR loci with consisting of tri- to hexa-nucleotide motifs, and 34 highly polymorphic primer pairs were selected to analyze the diversity of 231 germplasm resources. These primers amplified 219 alleles (Na) across 231 accessions, averaging of 6.441 alleles for per marker. The polymorphic information content (PIC) values ranged from 0.509 to 0.757 with a mean of 0.620. Genetic diversity varied among populations, with Southeast Asia showing the highest diversity, and Australia the lowest. Population structure analysis, divided the accessions into two groups, Group I (India) and Group II (Southeast Asia), containing 104 and 127 accessions, respectively, consistent with results from phylogenetic analysis and principal component analysis (PCA). Sixteen SSR primer pairs capable of distinguishing all tested accessions, were selected as core primers for constructing fingerprints of 229 mango accessions. These findings offer valuable resources for enhancing the utilization of mango germplasm in breeding programs.
Keywords: finger-printing; genetic diversity; genomic SSR; mango (Mangifera indica L.).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


References
-
- Tharanathan R.N., Yashoda H.M., Prabha T.N. Mango (Mangifera indica L.),“The king of fruits”—An overview. Food Rev. Int. 2006;22:95–123. doi: 10.1080/87559120600574493. - DOI
-
- Mukherjee S.K. Origin of mango (Mangifera indica) Econ. Bot. 1972;26:260–264. doi: 10.1007/BF02861039. - DOI
-
- Yadav D., Singh S.P. Mango: History origin and distribution. J. Pharmacogn. Phytochem. 2017;6:1257–1262.
-
- Litz R.E. The Mango: Botany, Production and Uses. CABI; Wallingford, UK: 2009.
MeSH terms
Substances
Grants and funding
- 32360727/National Natural Science Foundation of China
- CATASCXTD202405/Chinese Academy of Tropical Agricultural Sciences for Science and Technology Innovation Team of National Tropical Agricultural Science Center
- ZDYF2022XDNY255/Hainan Province Key Research and Development Plan
- 2022-NPY-00-030/Guangdong Province Seed Industry Revitalization Project
- CARS-31/China Agriculture Research System of MOF and MARA
LinkOut - more resources
Full Text Sources
Other Literature Sources

