MET Exon 14 Skipping and Novel Actionable Variants: Diagnostic and Therapeutic Implications in Latin American Non-Small-Cell Lung Cancer Patients
- PMID: 39769478
- PMCID: PMC11677537
- DOI: 10.3390/ijms252413715
MET Exon 14 Skipping and Novel Actionable Variants: Diagnostic and Therapeutic Implications in Latin American Non-Small-Cell Lung Cancer Patients
Abstract
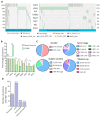
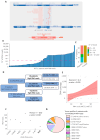
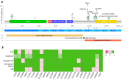
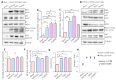
Targeted therapy indications for actionable variants in non-small-cell lung cancer (NSCLC) have primarily been studied in Caucasian populations, with limited data on Latin American patients. This study utilized a 52-genes next-generation sequencing (NGS) panel to analyze 1560 tumor biopsies from NSCLC patients in Chile, Brazil, and Peru. The RNA sequencing reads and DNA coverage were correlated to improve the detection of the actionable MET exon 14 skipping variant (METex14). The pathogenicity of MET variants of uncertain significance (VUSs) was assessed using bioinformatic methods, based on their predicted driver potential. The effects of the predicted drivers VUS T992I and H1094Y on c-MET signaling activation, proliferation, and migration were evaluated in HEK293T, BEAS-2B, and H1993 cell lines. Subsequently, c-Met inhibitors were tested in 2D and 3D cell cultures, and drug affinity was determined using 3D structure simulations. The prevalence of MET variants in the South American cohort was 8%, and RNA-based diagnosis detected 27% more cases of METex14 than DNA-based methods. Notably, 20% of METex14 cases with RNA reads below the detection threshold were confirmed using DNA analysis. The novel actionable T992I and H1094Y variants induced proliferation and migration through c-Met/Akt signaling. Both variants showed sensitivity to crizotinib and savolitinib, but the H1094Y variant exhibited reduced sensitivity to capmatinib. These findings highlight the importance of RNA-based METex14 diagnosis and reveal the drug sensitivity profiles of novel actionable MET variants from an understudied patient population.
Keywords: MET exon 14 skipping; c-Met inhibitors; next-generation sequencing; non-small-cell lung cancer; novel actionable variants.
Conflict of interest statement
R.A. received honoraria for conferences, advisory boards, and educational activities from Roche and Janssen, as well as grants and support for scientific research from Pfizer, Roche, and Thermo Fischer Scientific. The founders had no role in the study’s design; in the collection, analysis, or interpretation of data; the writing of the manuscript; or the decision to publish the results.
Figures

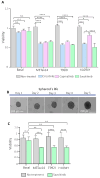

References
-
- Passiglia F., Bertolaccini L., Del Re M., Facchinetti F., Ferrara R., Franchina T., Malapelle U., Menis J., Passaro A., Pilotto S., et al. Diagnosis and treatment of early and locally advanced non-small-cell lung cancer: The 2019 AIOM (Italian Association of Medical Oncology) clinical practice guidelines. Crit. Rev. Oncol. Hematol. 2020;148:102862. doi: 10.1016/j.critrevonc.2019.102862. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

