The Effect of Oral Care Product Ingredients on Oral Pathogenic Bacteria Transcriptomics Through RNA-Seq
- PMID: 39770870
- PMCID: PMC11728304
- DOI: 10.3390/microorganisms12122668
The Effect of Oral Care Product Ingredients on Oral Pathogenic Bacteria Transcriptomics Through RNA-Seq
Abstract
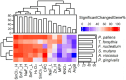
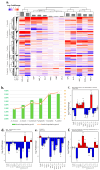
Various ingredients are utilized to inhibit the growth of harmful bacteria associated with cavities, gum disease, and bad breath. However, the precise mechanisms by which these ingredients affect the oral microbiome have not been fully understood at the molecular level. To elucidate the molecular mechanisms, a high-throughput bacterial transcriptomics study was conducted, and the gene expression profiles of six common oral bacteria, including two Gram-positive bacteria (Actinomyces viscosus, Streptococcus mutans) and four Gram-negative bacteria (Porphyromonas gingivalis, Tannerella forsythia, Fusobacterium nucleatum, and Prevotella pallens), were analyzed. The bacteria were exposed to nine common ingredients in toothpaste and mouthwash at different concentrations (stannous fluoride, stannous chloride, arginine bicarbonate, cetylpyridinium chloride, sodium monofluorophosphate, sodium fluoride, potassium nitrate, zinc phosphate, and hydrogen peroxide). Across 78 ingredient-microorganism pairs with 360 treatment-control combinations, significant and reproducible ingredient-based transcriptional response profiles were observed, providing valuable insights into the effects of these ingredients on the oral microbiome at the molecular level. This research shows that oral care product ingredients applied at biologically relevant concentrations manifest differential effects on the transcriptomics of bacterial genes in a variety of oral periodontal pathogenic bacteria. Stannous fluoride, stannous chloride, and cetylpyridinium chloride showed the most robust efficacy in inhibiting the growth or gene expression of various bacteria and pathogenic pathways. Combining multiple ingredients targeting different mechanisms might be more efficient than single ingredients in complex oral microbiomes.
Keywords: RNA-Seq; bacteria; cetylpyridinium chloride; oral care; pathogen; stannous fluoride; transcriptomics; virulence factors.
Conflict of interest statement
All authors are current employees of Procter & Gamble. The research cited in this document was funded by Procter & Gamble.
Figures





References
-
- WHO . Global Oral Health Status Report: Towards Universal Health Coverage for Oral Health by 2030. World Health Organization; Geneva, Switzerland: 2022. [(accessed on 8 November 2024)]. Available online: https://iris.who.int/handle/10665/364454.
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

