Sequence Analysis of microRNAs Encoded by Simian Lymphocryptoviruses
- PMID: 39772230
- PMCID: PMC11680086
- DOI: 10.3390/v16121923
Sequence Analysis of microRNAs Encoded by Simian Lymphocryptoviruses
Abstract
Lymphocryptoviruses (LCVs) are ubiquitous gamma-herpesviruses that establish life-long infections in both humans and non-human primates (NHPs). In immunocompromised hosts, LCV infections are commonly associated with B cell disorders and malignancies such as lymphoma. In this study, we evaluated simian LCV-encoded small microRNAs (miRNAs) present in lymphoblastoid cell lines (LCLs) derived from a Mauritian cynomolgus macaque (Macaca fascicularis) with cyLCV-associated post-transplant lymphoproliferative disease (PTLD) as well as the viral miRNAs expressed in a baboon (Papio hamadryas) LCL that harbors CeHV12. Via sequence comparisons, we further predicted viral miRNAs encoded by LCVs that infect two additional NHP species: stump-tailed macaques (Macaca arctoides) and bonobos (Pan paniscus). Together, these species represent two arms of the primate phylogeny: Hominoids (Pan) and Old-World monkeys (Macaca, Papio). Through our analysis, we defined sequences for >95 viral miRNAs encoded by these four NHP LCVs. Our study provides the most comprehensive annotation of NHP LCV miRNAs to date, yielding a resource for developing sequence-specific reagents to detect these molecules. Importantly, we further demonstrate that cyLCV miRNAs can be detected in circulation in vivo and have biomarker potential for LCV-related PTLD.
Keywords: biomarkers; herpesvirus; lymphocryptovirus; microRNAs.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
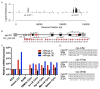



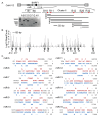

References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources

