CXCL12 ameliorates neutrophilia and disease severity in SARS-CoV-2 infection
- PMID: 39773555
- PMCID: PMC11827850
- DOI: 10.1172/JCI188222
CXCL12 ameliorates neutrophilia and disease severity in SARS-CoV-2 infection
Abstract
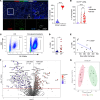
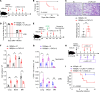
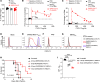
Neutrophils, particularly low-density neutrophils (LDNs), are believed to contribute to acute COVID-19 severity. Here, we showed that neutrophilia can be detected acutely and even months after SARS-CoV-2 infection in patients and mice, while neutrophil depletion reduced disease severity in mice. A key factor in neutrophilia and severe disease in infected mice was traced to the chemokine CXCL12 secreted by bone marrow cells and unexpectedly, endothelial cells. CXCL12 levels were negatively correlated with LDN numbers in longitudinal analyses of patient blood samples. CXCL12 blockade in SARS-CoV-2-infected mice increased blood/lung neutrophil numbers, thereby accelerating disease progression without changing lung virus titers. The exaggerated mortality caused by CXCL12 blockade could be reversed by neutrophil depletion. In addition, blocking interactions between SARS-CoV-2 and angiotensin-converting enzyme 2 (ACE2) reduced CXCL12 levels, suggesting a signal transduction from virus-mediated ACE2 ligation to increased CXCL12 secretion. Collectively, these results demonstrate a previously unappreciated role of CXCL12 in diminishing neutrophilia, including low-density neutrophilia, and its deleterious effects in SARS-CoV-2 infections. The results also support the involvement of SARS-CoV-2-endothelial cell interactions in viral pathogenesis.
Keywords: COVID-19; Chemokines; Endothelial cells; Infectious disease; Neutrophils.
Conflict of interest statement
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

