AozC, a zn(II)2Cys6 transcription factor, negatively regulates salt tolerance in Aspergillus oryzae by controlling fatty acid biosynthesis
- PMID: 39773712
- PMCID: PMC11706192
- DOI: 10.1186/s12934-024-02639-z
AozC, a zn(II)2Cys6 transcription factor, negatively regulates salt tolerance in Aspergillus oryzae by controlling fatty acid biosynthesis
Abstract
Background: In the soy sauce fermentation industry, Aspergillus oryzae (A. oryzae) plays an essential role and is frequently subjected to high salinity levels, which pose a significant osmotic stress. This environmental challenge necessitates the activation of stress response mechanisms within the fungus. The Zn(II)2Cys6 family of transcription factors, known for their zinc binuclear cluster-containing proteins, are key regulators in fungi, modulating various cellular functions such as stress adaptation and metabolic pathways.
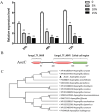
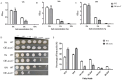
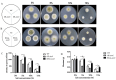
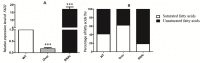
Results: Overexpression of AozC decreased growth rates in the presence of salt, while its knockdown enhanced growth, the number of spores, and biomass, particularly under conditions of 15% salt concentration, doubling these metrics compared to the wild type. Conversely, the knockdown of AozC via RNA interference significantly enhanced spore density and dry biomass, particularly under 15% salt stress, where these parameters were markedly improved over the wild type strain. Moreover, the overexpression of AozC led to a downregulation of the FAD2 gene, a pivotal enzyme in the biosynthesis of unsaturated fatty acids (UFAs), which are essential for preserving cell membrane fluidity and integrity under saline conditions. Transcriptome profiling further exposed the influence of AozC on the regulation of UFA biosynthesis and the modulation of critical stress response pathways. Notably, the regulatory role of AozC in the mitogen-activated protein kinase (MAPK) signaling and ABC transporters pathways was highlighted, underscoring its significance in cellular osmotic balance and endoplasmic reticulum homeostasis. These findings collectively indicate that AozC functions as a negative regulator of salt tolerance in A. oryzae.
Conclusion: This research suggest that AozC acts as a negative regulator in salt tolerance and modulates fatty acid biosynthesis in response to osmotic stress. These results provide insights into the regulatory mechanisms of stress adaptation in A. oryzae.
Keywords: Aspergillus oryzae; Differentially expressed genes; Fatty acid; Salt treatment; Transcription factor; Transcriptome; Unsaturated fatty acids.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures
Similar articles
-
Genome-wide identification and expression profile analysis of the HOG gene family in Aspergillus oryzae.World J Microbiol Biotechnol. 2018 Feb 9;34(2):35. doi: 10.1007/s11274-018-2419-6. World J Microbiol Biotechnol. 2018. PMID: 29427255
-
Deep sequencing analysis of transcriptomes in Aspergillus oryzae in response to salinity stress.Appl Microbiol Biotechnol. 2018 Jan;102(2):897-906. doi: 10.1007/s00253-017-8603-z. Epub 2017 Nov 3. Appl Microbiol Biotechnol. 2018. PMID: 29101425
-
Heterologous expression of AoD9D enhances salt tolerance with increased accumulation of unsaturated fatty acid in transgenic Saccharomyces cerevisiae.J Ind Microbiol Biotechnol. 2019 Feb;46(2):231-239. doi: 10.1007/s10295-018-02123-9. Epub 2019 Jan 2. J Ind Microbiol Biotechnol. 2019. PMID: 30604237
-
Transcriptional and post-transcriptional regulation of genes encoding secretory proteins in Aspergillus oryzae.Biosci Biotechnol Biochem. 2024 Mar 22;88(4):381-388. doi: 10.1093/bbb/zbae004. Biosci Biotechnol Biochem. 2024. PMID: 38211972 Review.
-
The Translation Initiation Factor eIF2Bα Regulates Development, Stress Response, Amylase Production, and Kojic Acid Synthesis in the Fungus Aspergillus oryzae.Curr Microbiol. 2025 Jan 5;82(2):70. doi: 10.1007/s00284-024-04051-7. Curr Microbiol. 2025. PMID: 39756002 Review.
References
-
- Liu J, Li J, Shin HD, Du G, Chen J, Liu L. Metabolic engineering of aspergillus oryzae for efficient production of l-malate directly from corn starch. J Biotechnol. 2017;262:40–6. 10.1016/j.jbiotec.2017.09.021. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

